Probe CUST_3166_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3166_PI426222305 | JHI_St_60k_v1 | DMT400000365 | GATTGAACTTCAGAAAATACCATTTCATGATGAAACATGAAACGCCCATTCAGGATTTTG |
All Microarray Probes Designed to Gene DMG400000129
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3166_PI426222305 | JHI_St_60k_v1 | DMT400000365 | GATTGAACTTCAGAAAATACCATTTCATGATGAAACATGAAACGCCCATTCAGGATTTTG |
Microarray Signals from CUST_3166_PI426222305
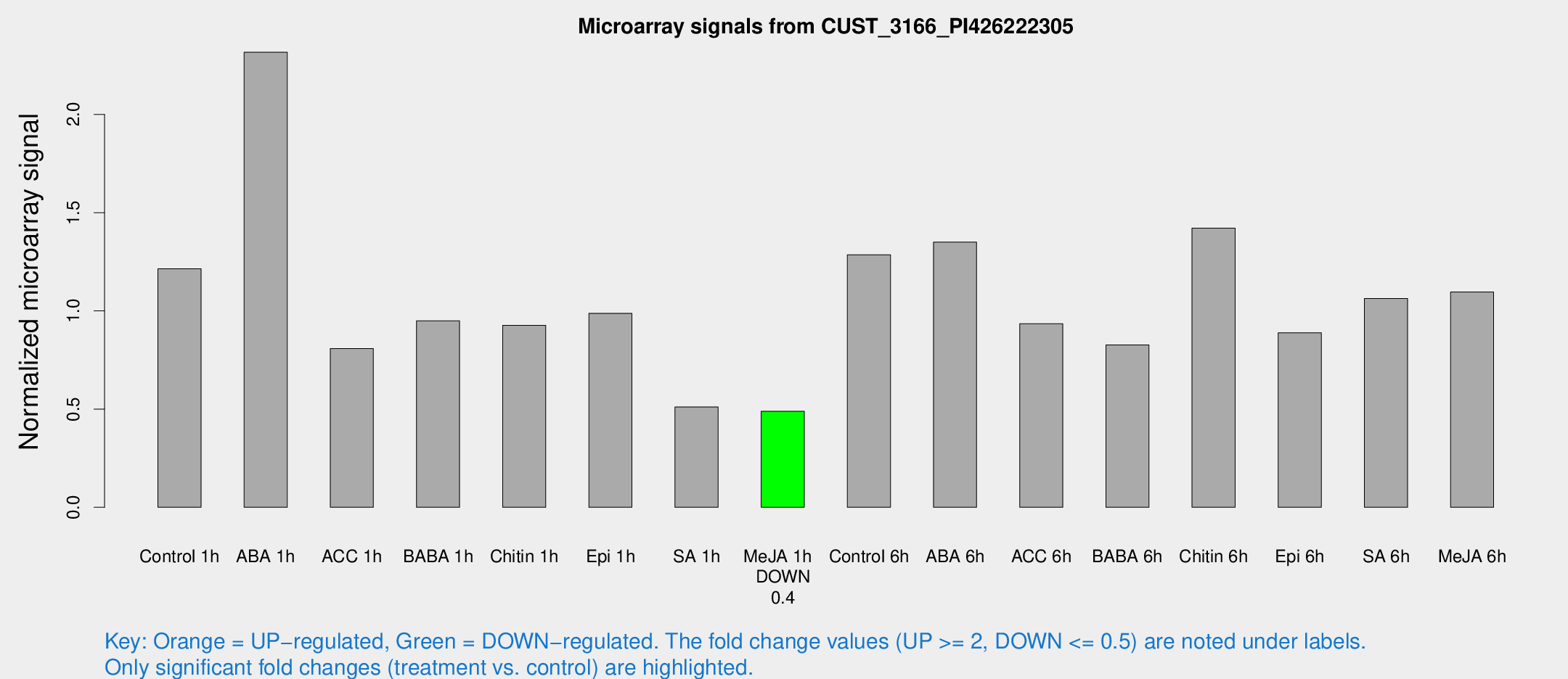
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 90.1488 | 12.6093 | 1.21427 | 0.135237 |
| ABA 1h | 152.078 | 20.8573 | 2.31693 | 0.220985 |
| ACC 1h | 61.7995 | 9.40151 | 0.808245 | 0.0783536 |
| BABA 1h | 71.0305 | 16.5406 | 0.949293 | 0.160506 |
| Chitin 1h | 60.8053 | 4.68803 | 0.92669 | 0.0715649 |
| Epi 1h | 63.1357 | 8.39236 | 0.987519 | 0.102004 |
| SA 1h | 43.8547 | 13.5668 | 0.510505 | 0.201241 |
| Me-JA 1h | 29.6359 | 4.4735 | 0.488957 | 0.0603802 |
| Control 6h | 94.2288 | 7.64341 | 1.28597 | 0.11753 |
| ABA 6h | 107.352 | 17.1057 | 1.35058 | 0.190762 |
| ACC 6h | 78.8412 | 8.89187 | 0.934727 | 0.193995 |
| BABA 6h | 70.8259 | 15.8739 | 0.826601 | 0.174864 |
| Chitin 6h | 112.729 | 16.9482 | 1.42105 | 0.233079 |
| Epi 6h | 73.1637 | 5.59406 | 0.888299 | 0.0967464 |
| SA 6h | 76.469 | 5.51907 | 1.06282 | 0.0924845 |
| Me-JA 6h | 87.704 | 29.0396 | 1.09602 | 0.267994 |
Source Transcript PGSC0003DMT400000365 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G46690.1 | +1 | 7e-33 | 120 | 67/111 (60%) | SAUR-like auxin-responsive protein family | chr2:19180904-19181269 FORWARD LENGTH=121 |
