Probe CUST_31591_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31591_PI426222305 | JHI_St_60k_v1 | DMT400042521 | TGATCTATGCTATTTTCTTAGCTCCCTTGCTACTATGGTCATGCGATGCATCTCGAGAAC |
All Microarray Probes Designed to Gene DMG400016493
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31591_PI426222305 | JHI_St_60k_v1 | DMT400042521 | TGATCTATGCTATTTTCTTAGCTCCCTTGCTACTATGGTCATGCGATGCATCTCGAGAAC |
| CUST_31583_PI426222305 | JHI_St_60k_v1 | DMT400042522 | TCTGTATGCAGAACTCGCGTCAAGCAAGTTTCCACCACGTGTTGTATGTCTTCATTGTGT |
Microarray Signals from CUST_31591_PI426222305
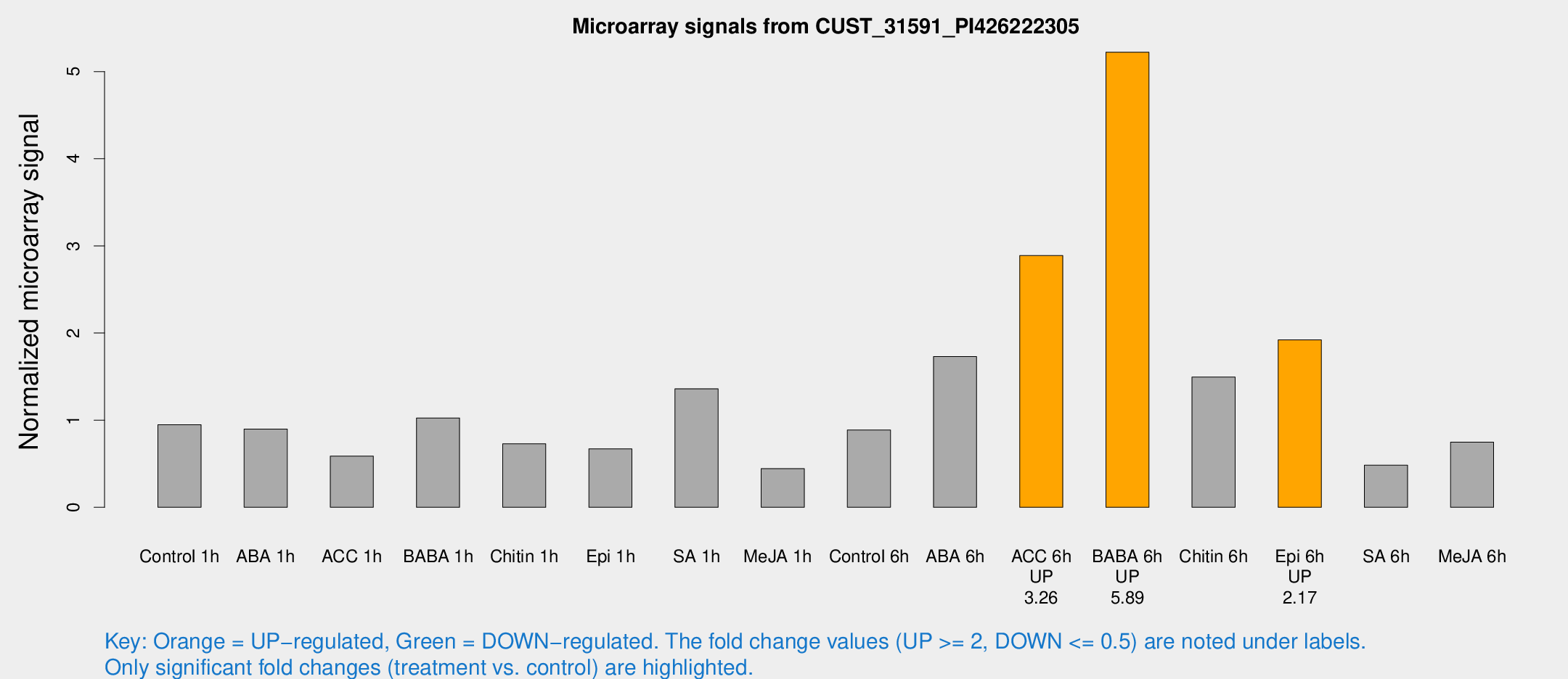
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 198.555 | 68.4345 | 0.947321 | 0.261212 |
| ABA 1h | 153.312 | 22.5544 | 0.897994 | 0.101672 |
| ACC 1h | 138.839 | 49.4364 | 0.586913 | 0.284171 |
| BABA 1h | 186.596 | 11.2853 | 1.02509 | 0.0618835 |
| Chitin 1h | 132.451 | 35.8488 | 0.728033 | 0.171939 |
| Epi 1h | 114.478 | 22.9125 | 0.671166 | 0.146261 |
| SA 1h | 274.466 | 54.6531 | 1.35966 | 0.345701 |
| Me-JA 1h | 70.9672 | 13.3168 | 0.443115 | 0.096936 |
| Control 6h | 170.415 | 23.1941 | 0.887187 | 0.113726 |
| ABA 6h | 353.05 | 50.4157 | 1.72908 | 0.146148 |
| ACC 6h | 660.065 | 159.328 | 2.88911 | 0.951561 |
| BABA 6h | 1137.69 | 200.497 | 5.22203 | 1.13575 |
| Chitin 6h | 300.342 | 17.7467 | 1.4941 | 0.0880754 |
| Epi 6h | 413.869 | 46.7008 | 1.92221 | 0.288949 |
| SA 6h | 93.811 | 19.7641 | 0.482746 | 0.0659288 |
| Me-JA 6h | 186.399 | 100.185 | 0.747197 | 0.389425 |
Source Transcript PGSC0003DMT400042521 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
