Probe CUST_31543_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31543_PI426222305 | JHI_St_60k_v1 | DMT400073330 | AAATACTACGTCTCAACGCAATCCACCAGGAGAATACAAATATGCAGTACCGGTTTGGGA |
All Microarray Probes Designed to Gene DMG400028498
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31543_PI426222305 | JHI_St_60k_v1 | DMT400073330 | AAATACTACGTCTCAACGCAATCCACCAGGAGAATACAAATATGCAGTACCGGTTTGGGA |
Microarray Signals from CUST_31543_PI426222305
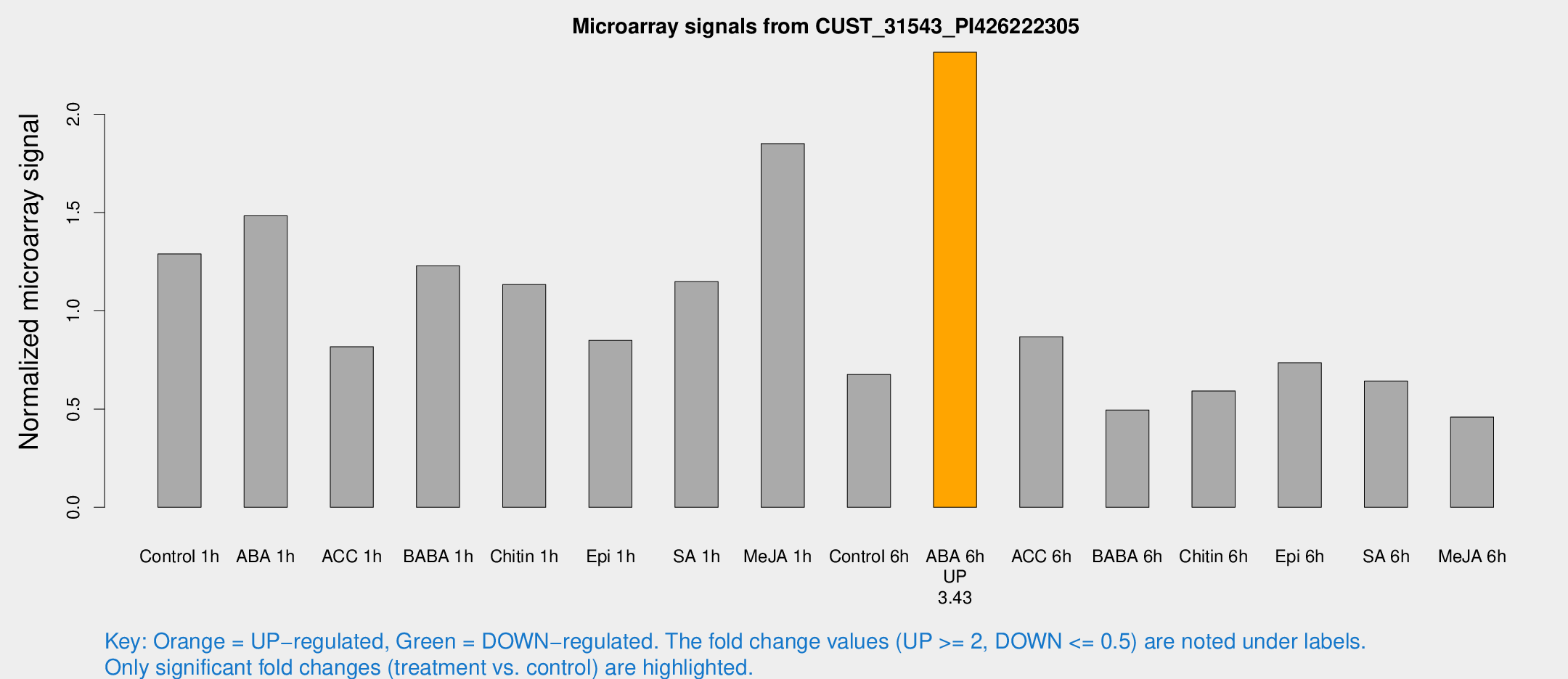
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 45.7981 | 5.91629 | 1.28923 | 0.239876 |
| ABA 1h | 46.3331 | 4.88446 | 1.48364 | 0.147729 |
| ACC 1h | 35.7789 | 12.4931 | 0.817128 | 0.393843 |
| BABA 1h | 41.8426 | 4.58632 | 1.22903 | 0.260964 |
| Chitin 1h | 36.1992 | 4.65385 | 1.1335 | 0.136517 |
| Epi 1h | 26.8488 | 5.86967 | 0.849457 | 0.186364 |
| SA 1h | 41.6567 | 4.33064 | 1.1485 | 0.178582 |
| Me-JA 1h | 55.2968 | 10.9344 | 1.85073 | 0.613151 |
| Control 6h | 26.8291 | 8.46827 | 0.67561 | 0.201847 |
| ABA 6h | 86.6947 | 7.78179 | 2.31602 | 0.17029 |
| ACC 6h | 34.8681 | 5.05922 | 0.867562 | 0.123349 |
| BABA 6h | 23.3145 | 9.53379 | 0.495437 | 0.226517 |
| Chitin 6h | 23.2094 | 5.11248 | 0.592331 | 0.166755 |
| Epi 6h | 29.7483 | 4.97625 | 0.735857 | 0.130706 |
| SA 6h | 24.6529 | 7.83443 | 0.642471 | 0.247638 |
| Me-JA 6h | 17.527 | 4.84083 | 0.459137 | 0.194969 |
Source Transcript PGSC0003DMT400073330 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G10970.1 | +2 | 1e-20 | 89 | 59/129 (46%) | C2H2 and C2HC zinc fingers superfamily protein | chr5:3469268-3470086 FORWARD LENGTH=272 |
