Probe CUST_31537_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31537_PI426222305 | JHI_St_60k_v1 | DMT400042632 | AAGAGGTTTTCTTACCTAGAGAACTTAGGTATGTGTAGAAGTCCAACTGCACGTCACTAA |
All Microarray Probes Designed to Gene DMG400016533
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31537_PI426222305 | JHI_St_60k_v1 | DMT400042632 | AAGAGGTTTTCTTACCTAGAGAACTTAGGTATGTGTAGAAGTCCAACTGCACGTCACTAA |
Microarray Signals from CUST_31537_PI426222305
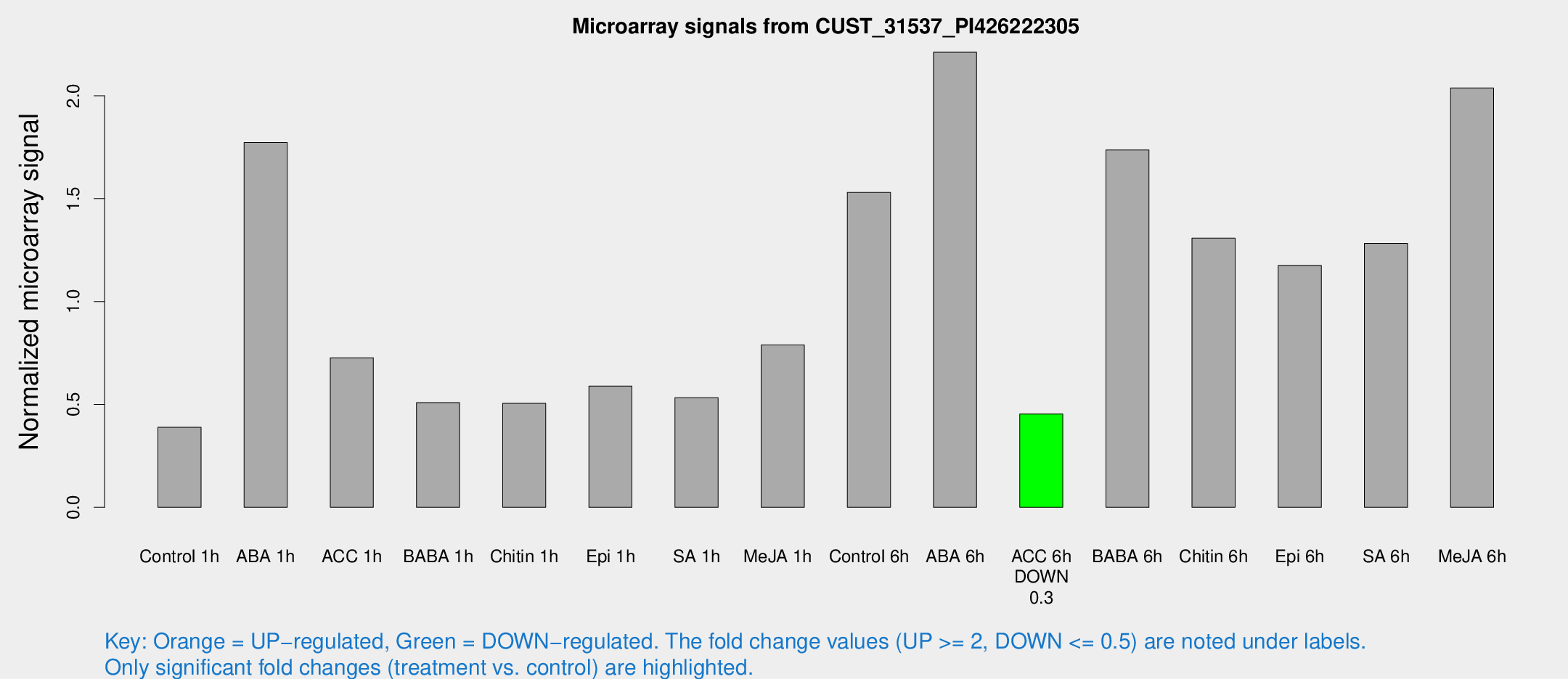
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.61158 | 3.69536 | 0.388707 | 0.169339 |
| ABA 1h | 40.7411 | 11.7889 | 1.77289 | 0.504327 |
| ACC 1h | 18.2119 | 4.17427 | 0.726647 | 0.201058 |
| BABA 1h | 12.5793 | 3.87015 | 0.508576 | 0.194345 |
| Chitin 1h | 11.6488 | 3.71056 | 0.504918 | 0.19396 |
| Epi 1h | 11.9942 | 3.61478 | 0.588682 | 0.178221 |
| SA 1h | 14.3662 | 4.21315 | 0.532901 | 0.254141 |
| Me-JA 1h | 15.2402 | 3.83761 | 0.789074 | 0.202719 |
| Control 6h | 37.6725 | 7.92604 | 1.53002 | 0.200733 |
| ABA 6h | 55.8369 | 6.27661 | 2.21167 | 0.199755 |
| ACC 6h | 12.2808 | 4.67039 | 0.453507 | 0.178446 |
| BABA 6h | 45.7177 | 4.85559 | 1.73682 | 0.186234 |
| Chitin 6h | 33.873 | 6.03564 | 1.30824 | 0.302709 |
| Epi 6h | 31.1033 | 4.78017 | 1.17558 | 0.180661 |
| SA 6h | 30.2038 | 4.29749 | 1.28274 | 0.191168 |
| Me-JA 6h | 49.2258 | 8.71722 | 2.03775 | 0.208224 |
Source Transcript PGSC0003DMT400042632 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G79860.1 | +1 | 0.0 | 555 | 296/446 (66%) | RHO guanyl-nucleotide exchange factor 12 | chr1:30042172-30044092 REVERSE LENGTH=515 |
