Probe CUST_31536_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31536_PI426222305 | JHI_St_60k_v1 | DMT400042531 | CGATTTATCGACTCTTATGGCTCAATTGCCAATCAAGAAAGGACTTTCGGAGTATTATGA |
All Microarray Probes Designed to Gene DMG400016497
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31536_PI426222305 | JHI_St_60k_v1 | DMT400042531 | CGATTTATCGACTCTTATGGCTCAATTGCCAATCAAGAAAGGACTTTCGGAGTATTATGA |
| CUST_31449_PI426222305 | JHI_St_60k_v1 | DMT400042530 | TTTGTGAATGTAGGAAAGGACTTTCGGAGTATTATGATGGGAAATCGGAGTCATTTACGT |
Microarray Signals from CUST_31536_PI426222305
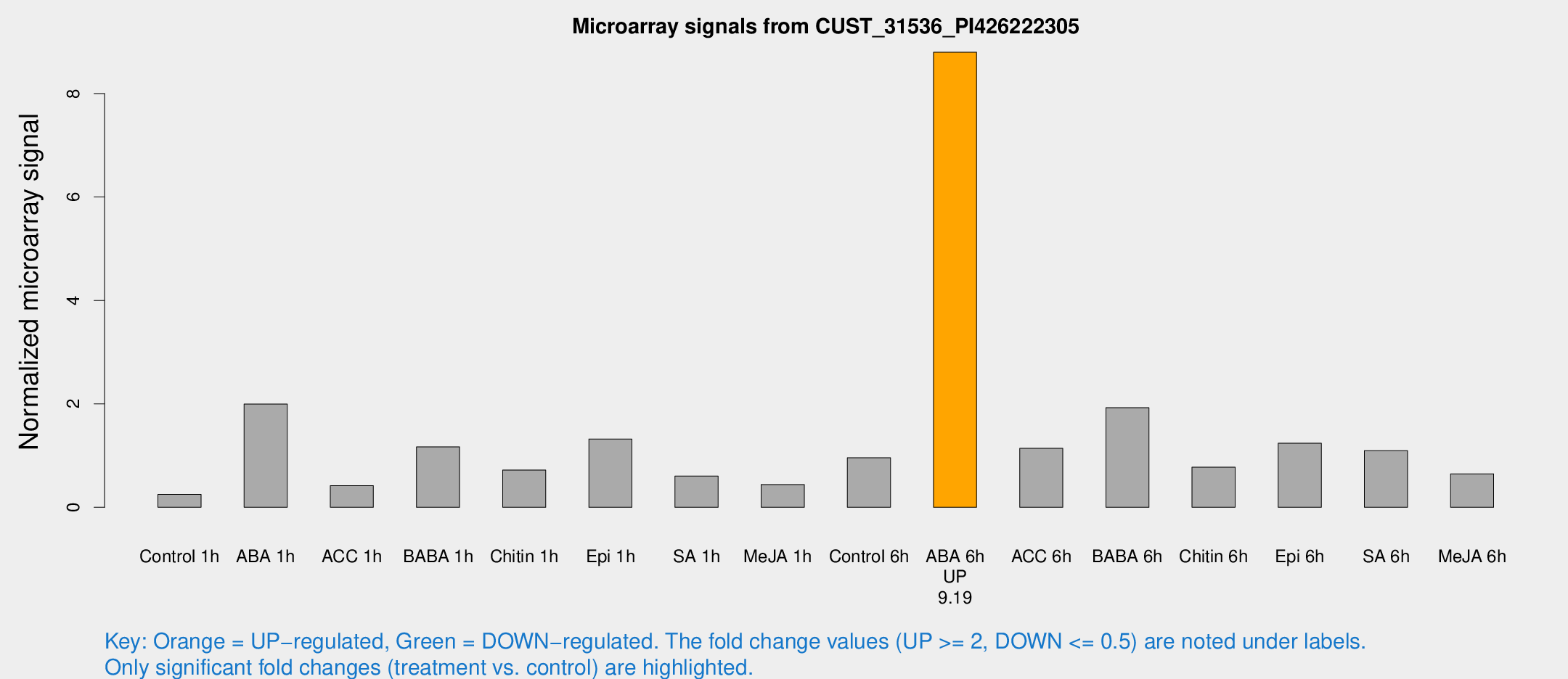
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 10.2788 | 3.2123 | 0.249642 | 0.0779622 |
| ABA 1h | 102.49 | 60.3103 | 1.99544 | 1.23523 |
| ACC 1h | 21.9847 | 8.48273 | 0.418233 | 0.209345 |
| BABA 1h | 54.3745 | 18.6589 | 1.16893 | 0.413054 |
| Chitin 1h | 28.5087 | 7.46123 | 0.720357 | 0.161993 |
| Epi 1h | 55.7895 | 22.2664 | 1.31967 | 0.590868 |
| SA 1h | 30.0891 | 10.3774 | 0.604564 | 0.29755 |
| Me-JA 1h | 18.8802 | 7.50592 | 0.441148 | 0.241521 |
| Control 6h | 41.3481 | 9.20644 | 0.95787 | 0.193292 |
| ABA 6h | 406.856 | 97.2985 | 8.8001 | 1.58584 |
| ACC 6h | 55.6066 | 9.88519 | 1.13935 | 0.14214 |
| BABA 6h | 96.1695 | 28.5112 | 1.92628 | 0.506304 |
| Chitin 6h | 38.5629 | 13.8624 | 0.774986 | 0.281913 |
| Epi 6h | 58.2777 | 6.42625 | 1.23973 | 0.109229 |
| SA 6h | 47.4031 | 11.4168 | 1.09666 | 0.16923 |
| Me-JA 6h | 41.7666 | 19.1607 | 0.646813 | 0.751853 |
Source Transcript PGSC0003DMT400042531 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G56550.1 | +2 | 2e-10 | 59 | 37/86 (43%) | oxidative stress 3 | chr5:22895847-22896502 REVERSE LENGTH=172 |
