Probe CUST_31043_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31043_PI426222305 | JHI_St_60k_v1 | DMT400010515 | CTCCAACTAAGCTACACGTTCTGGACTAATCTCTAAAGAGAAATATTACTATGGTTTCTT |
All Microarray Probes Designed to Gene DMG400004109
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31043_PI426222305 | JHI_St_60k_v1 | DMT400010515 | CTCCAACTAAGCTACACGTTCTGGACTAATCTCTAAAGAGAAATATTACTATGGTTTCTT |
| CUST_31027_PI426222305 | JHI_St_60k_v1 | DMT400010514 | GACTATACCATCCTTTGGAACAATCATCATATAGTGTTCTGGACTAATCTCTAAAGAGAA |
Microarray Signals from CUST_31043_PI426222305
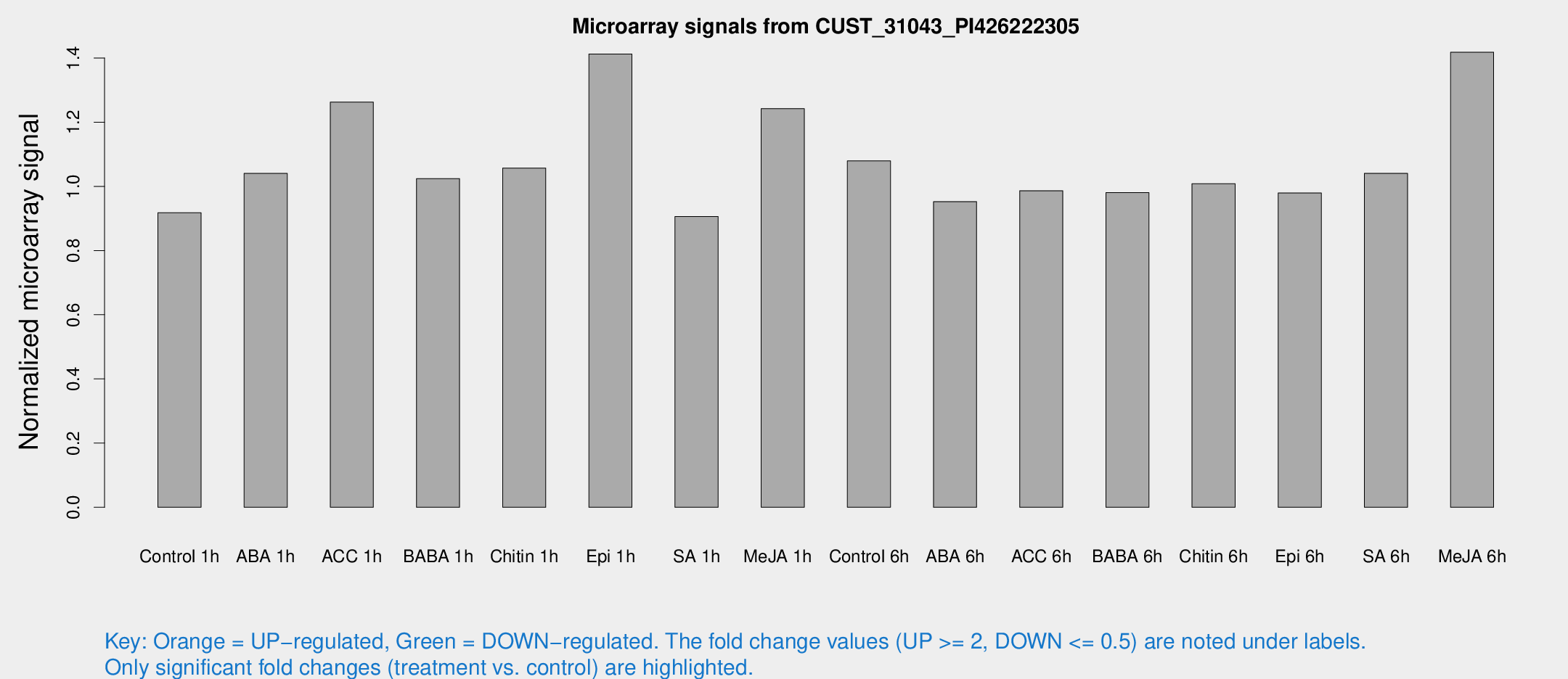
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.24489 | 3.04281 | 0.918032 | 0.532313 |
| ABA 1h | 5.25705 | 2.98141 | 1.04039 | 0.589158 |
| ACC 1h | 7.61286 | 3.3403 | 1.26269 | 0.587473 |
| BABA 1h | 5.64283 | 3.26018 | 1.02413 | 0.591463 |
| Chitin 1h | 5.35394 | 3.1151 | 1.05713 | 0.602187 |
| Epi 1h | 7.42696 | 3.11095 | 1.41256 | 0.67943 |
| SA 1h | 5.31689 | 3.0851 | 0.906225 | 0.525784 |
| Me-JA 1h | 5.86157 | 3.17875 | 1.24241 | 0.681108 |
| Control 6h | 6.23745 | 3.25558 | 1.07958 | 0.574352 |
| ABA 6h | 5.78868 | 3.35547 | 0.952467 | 0.551578 |
| ACC 6h | 6.61661 | 3.92815 | 0.986442 | 0.571309 |
| BABA 6h | 6.28268 | 3.6496 | 0.980425 | 0.568072 |
| Chitin 6h | 6.12931 | 3.55003 | 1.00814 | 0.583852 |
| Epi 6h | 6.35553 | 3.71019 | 0.979227 | 0.567049 |
| SA 6h | 5.8822 | 3.41221 | 1.04061 | 0.603256 |
| Me-JA 6h | 9.40107 | 3.86466 | 1.41793 | 0.661963 |
Source Transcript PGSC0003DMT400010515 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G10550.1 | +3 | 2e-65 | 209 | 93/165 (56%) | xyloglucan:xyloglucosyl transferase 33 | chr1:3479257-3480957 REVERSE LENGTH=310 |
