Probe CUST_31027_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31027_PI426222305 | JHI_St_60k_v1 | DMT400010514 | GACTATACCATCCTTTGGAACAATCATCATATAGTGTTCTGGACTAATCTCTAAAGAGAA |
All Microarray Probes Designed to Gene DMG400004109
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31043_PI426222305 | JHI_St_60k_v1 | DMT400010515 | CTCCAACTAAGCTACACGTTCTGGACTAATCTCTAAAGAGAAATATTACTATGGTTTCTT |
| CUST_31027_PI426222305 | JHI_St_60k_v1 | DMT400010514 | GACTATACCATCCTTTGGAACAATCATCATATAGTGTTCTGGACTAATCTCTAAAGAGAA |
Microarray Signals from CUST_31027_PI426222305
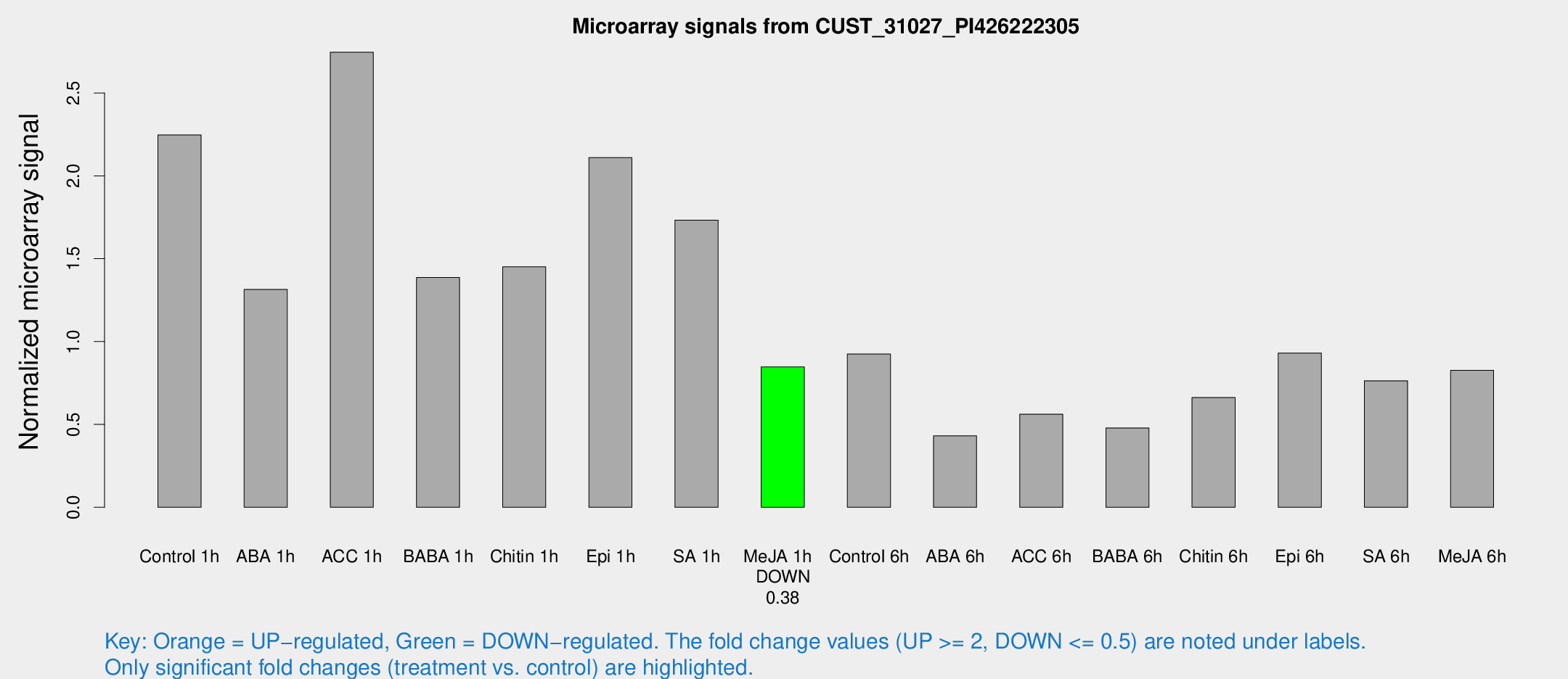
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 73.6748 | 16.5957 | 2.24677 | 0.344097 |
| ABA 1h | 36.6931 | 4.14224 | 1.31474 | 0.150308 |
| ACC 1h | 89.259 | 9.4919 | 2.74571 | 0.20481 |
| BABA 1h | 46.586 | 13.9133 | 1.38673 | 0.35539 |
| Chitin 1h | 43.804 | 10.2922 | 1.45074 | 0.330442 |
| Epi 1h | 57.5276 | 4.90784 | 2.11017 | 0.222768 |
| SA 1h | 61.7814 | 18.2541 | 1.73282 | 0.501009 |
| Me-JA 1h | 23.2634 | 5.59129 | 0.846923 | 0.195795 |
| Control 6h | 35.0973 | 13.4443 | 0.924779 | 0.396471 |
| ABA 6h | 14.8369 | 4.17278 | 0.43095 | 0.130125 |
| ACC 6h | 21.7828 | 5.30389 | 0.562106 | 0.138165 |
| BABA 6h | 19.5154 | 7.49031 | 0.478876 | 0.218063 |
| Chitin 6h | 22.9402 | 4.67256 | 0.662711 | 0.151024 |
| Epi 6h | 36.3729 | 12.1752 | 0.930902 | 0.299906 |
| SA 6h | 24.4004 | 4.40191 | 0.763093 | 0.148377 |
| Me-JA 6h | 28.5543 | 7.89397 | 0.826492 | 0.238137 |
Source Transcript PGSC0003DMT400010514 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G10550.1 | +3 | 9e-129 | 377 | 172/304 (57%) | xyloglucan:xyloglucosyl transferase 33 | chr1:3479257-3480957 REVERSE LENGTH=310 |
