Probe CUST_30940_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30940_PI426222305 | JHI_St_60k_v1 | DMT400040294 | TGCTACATGAAGTACAAGAAATCCTTGATTGCCTTTGTCTAGATGAAAATGAGCCAATTC |
All Microarray Probes Designed to Gene DMG400015601
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30995_PI426222305 | JHI_St_60k_v1 | DMT400040295 | GGTAGGTAAAATAATTCAACCTCATGTTATATTAGTACCATACCCAAGTCAAGGTCACAT |
| CUST_30940_PI426222305 | JHI_St_60k_v1 | DMT400040294 | TGCTACATGAAGTACAAGAAATCCTTGATTGCCTTTGTCTAGATGAAAATGAGCCAATTC |
Microarray Signals from CUST_30940_PI426222305
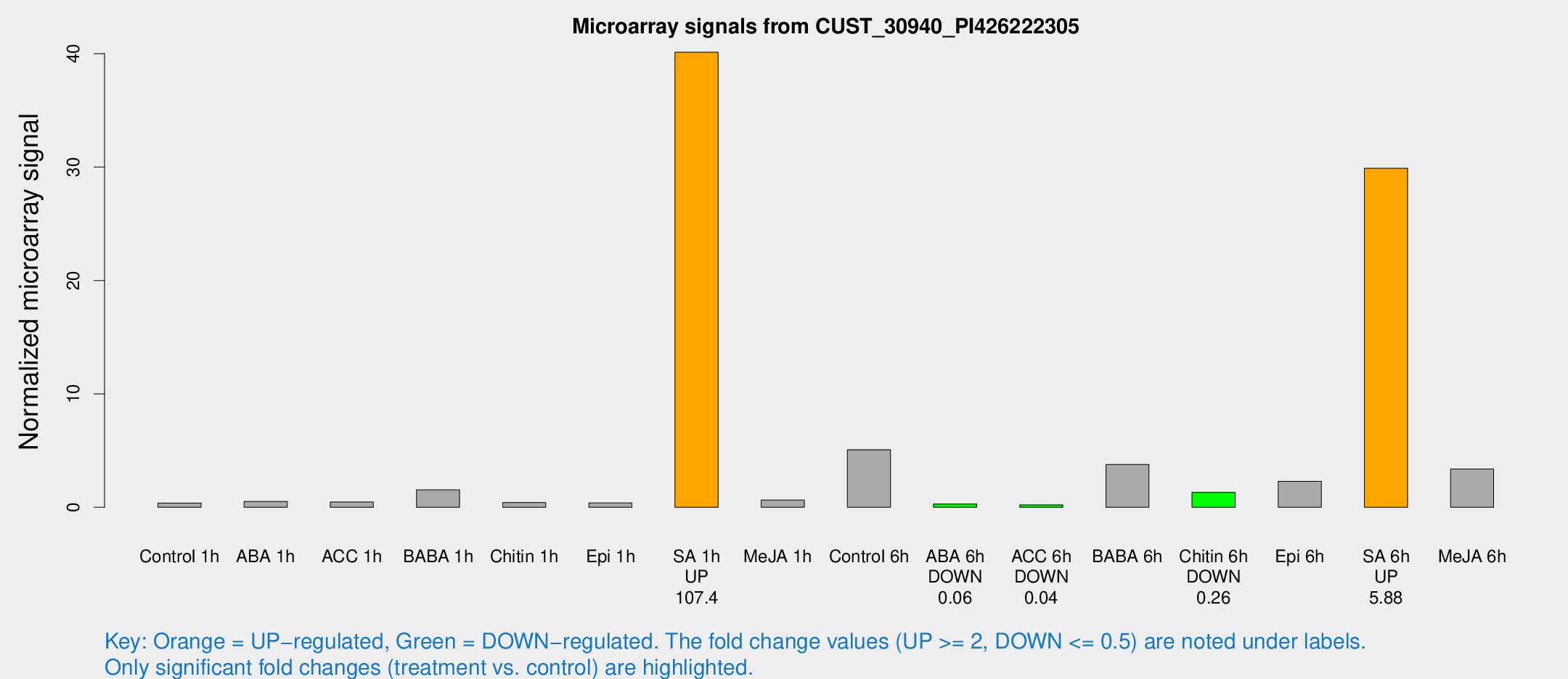
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 13.8053 | 7.18064 | 0.373644 | 0.186454 |
| ABA 1h | 19.3062 | 11.93 | 0.517989 | 0.422241 |
| ACC 1h | 16.7311 | 7.87898 | 0.466154 | 0.218118 |
| BABA 1h | 48.8596 | 18.9718 | 1.54042 | 0.873508 |
| Chitin 1h | 11.2371 | 3.39338 | 0.428385 | 0.139014 |
| Epi 1h | 11.4306 | 4.9957 | 0.387833 | 0.191795 |
| SA 1h | 1195.58 | 159.986 | 40.1294 | 3.30612 |
| Me-JA 1h | 15.5134 | 3.5527 | 0.644049 | 0.160012 |
| Control 6h | 176.235 | 78.6758 | 5.07937 | 2.00008 |
| ABA 6h | 8.99807 | 3.60068 | 0.287015 | 0.12457 |
| ACC 6h | 7.1433 | 4.25253 | 0.212786 | 0.123281 |
| BABA 6h | 124.333 | 20.5601 | 3.78217 | 0.760806 |
| Chitin 6h | 41.4234 | 7.27629 | 1.3215 | 0.224373 |
| Epi 6h | 85.5171 | 30.947 | 2.29187 | 1.29861 |
| SA 6h | 969.863 | 351.045 | 29.8875 | 9.06806 |
| Me-JA 6h | 109.122 | 33.8428 | 3.37759 | 1.46605 |
Source Transcript PGSC0003DMT400040294 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G24100.1 | +1 | 2e-106 | 320 | 153/264 (58%) | UDP-glucosyl transferase 74B1 | chr1:8525547-8527010 REVERSE LENGTH=460 |
