Probe CUST_30592_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30592_PI426222305 | JHI_St_60k_v1 | DMT400007974 | TATATCTGGTTCATATTGCTCTTGGCAGGGGGACAAAGGAAGGTGTCATTCTTTCCTTGG |
All Microarray Probes Designed to Gene DMG400003083
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30592_PI426222305 | JHI_St_60k_v1 | DMT400007974 | TATATCTGGTTCATATTGCTCTTGGCAGGGGGACAAAGGAAGGTGTCATTCTTTCCTTGG |
| CUST_30636_PI426222305 | JHI_St_60k_v1 | DMT400007975 | TAGGATTTCTCAGCACAAATTGCAGCAAAGCTCCAGTTTTCTTGGAAATAACCCTCTCAG |
Microarray Signals from CUST_30592_PI426222305
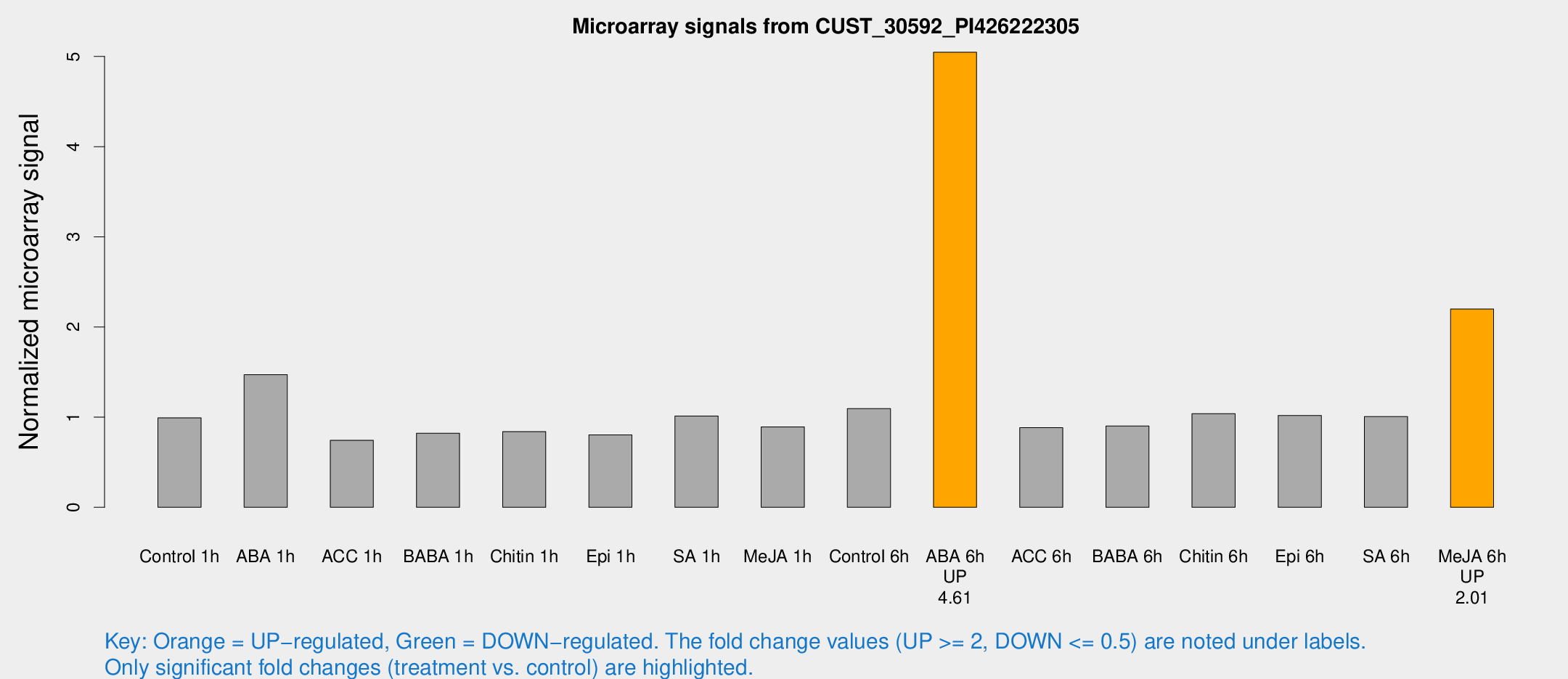
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2656.81 | 187.297 | 0.991604 | 0.0572664 |
| ABA 1h | 3529.32 | 434.882 | 1.47186 | 0.122752 |
| ACC 1h | 2241.99 | 601.188 | 0.743146 | 0.19955 |
| BABA 1h | 2224.09 | 461.814 | 0.822208 | 0.109997 |
| Chitin 1h | 2095.21 | 444.476 | 0.838643 | 0.124305 |
| Epi 1h | 1907.19 | 306.203 | 0.803582 | 0.134111 |
| SA 1h | 2830.7 | 411.29 | 1.01175 | 0.127077 |
| Me-JA 1h | 1970.12 | 239.169 | 0.892286 | 0.103943 |
| Control 6h | 3013.31 | 491.245 | 1.09469 | 0.102991 |
| ABA 6h | 14780.9 | 2519.86 | 5.04703 | 0.698355 |
| ACC 6h | 2708.31 | 157.299 | 0.882706 | 0.0915429 |
| BABA 6h | 2729.55 | 331.065 | 0.901271 | 0.129561 |
| Chitin 6h | 2953.77 | 170.969 | 1.03868 | 0.0829425 |
| Epi 6h | 3086.87 | 294.021 | 1.01762 | 0.0742528 |
| SA 6h | 2819.65 | 646.923 | 1.0061 | 0.146126 |
| Me-JA 6h | 5993.15 | 991.231 | 2.19853 | 0.310272 |
Source Transcript PGSC0003DMT400007974 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G29395.1 | +1 | 5e-16 | 80 | 51/107 (48%) | COLD REGULATED 314 INNER MEMBRANE 1 | chr1:10288345-10289539 REVERSE LENGTH=225 |
