Probe CUST_30347_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30347_PI426222305 | JHI_St_60k_v1 | DMT400069836 | AGGAGAAGTAGCTGTCCGATTTTTTATTGGTCTTGTTAAAATCCTACCTGCCAAATATGT |
All Microarray Probes Designed to Gene DMG400027154
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30339_PI426222305 | JHI_St_60k_v1 | DMT400069837 | GCCATCTTATGTTGTGTTGGTAAGCTGATGTTGAATGTTGAAAAAGGAGTAATAGATACT |
| CUST_30347_PI426222305 | JHI_St_60k_v1 | DMT400069836 | AGGAGAAGTAGCTGTCCGATTTTTTATTGGTCTTGTTAAAATCCTACCTGCCAAATATGT |
Microarray Signals from CUST_30347_PI426222305
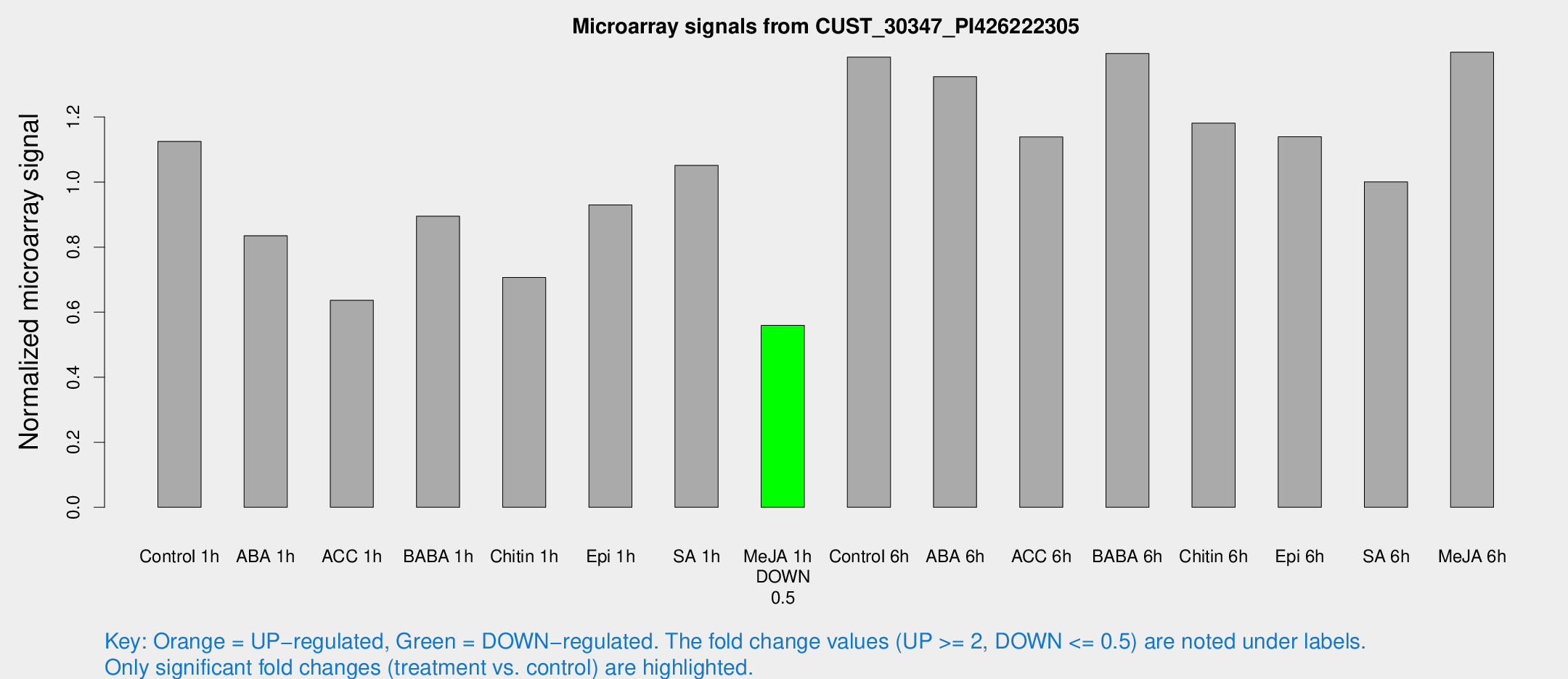
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 62.2566 | 6.86279 | 1.12501 | 0.0869616 |
| ABA 1h | 41.6004 | 6.80484 | 0.835052 | 0.0980152 |
| ACC 1h | 39.2913 | 10.5571 | 0.636473 | 0.166251 |
| BABA 1h | 47.5065 | 4.27788 | 0.895 | 0.0814923 |
| Chitin 1h | 35.3499 | 4.87525 | 0.706613 | 0.0755606 |
| Epi 1h | 44.9492 | 6.44687 | 0.92995 | 0.135416 |
| SA 1h | 59.0372 | 4.59215 | 1.05119 | 0.0818178 |
| Me-JA 1h | 25.2733 | 3.51027 | 0.559298 | 0.07884 |
| Control 6h | 94.0685 | 38.7169 | 1.38437 | 0.653576 |
| ABA 6h | 78.0416 | 8.94114 | 1.32428 | 0.0991506 |
| ACC 6h | 74.703 | 15.8015 | 1.13885 | 0.110915 |
| BABA 6h | 86.4648 | 9.63739 | 1.39553 | 0.123766 |
| Chitin 6h | 69.1525 | 5.56492 | 1.18144 | 0.092404 |
| Epi 6h | 77.096 | 24.8628 | 1.13939 | 0.359855 |
| SA 6h | 65.2659 | 24.5079 | 1.00088 | 0.385913 |
| Me-JA 6h | 81.6601 | 19.1985 | 1.39932 | 0.286992 |
Source Transcript PGSC0003DMT400069836 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G06440.1 | +1 | 2e-173 | 510 | 302/658 (46%) | Galactosyltransferase family protein | chr3:1972913-1975272 REVERSE LENGTH=619 |
