Probe CUST_30341_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30341_PI426222305 | JHI_St_60k_v1 | DMT400069793 | GGACCATTCTACTTGCCCTCCCAAACTCCCTCCATATTTTAGTACTTTTTATGGTTTGAT |
All Microarray Probes Designed to Gene DMG400027140
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30341_PI426222305 | JHI_St_60k_v1 | DMT400069793 | GGACCATTCTACTTGCCCTCCCAAACTCCCTCCATATTTTAGTACTTTTTATGGTTTGAT |
| CUST_30309_PI426222305 | JHI_St_60k_v1 | DMT400069794 | GGACCATTCTACTTGCCCTCCCAAACTCCCTCCATATTTTAGTACTTTTTATGGTTTGAT |
Microarray Signals from CUST_30341_PI426222305
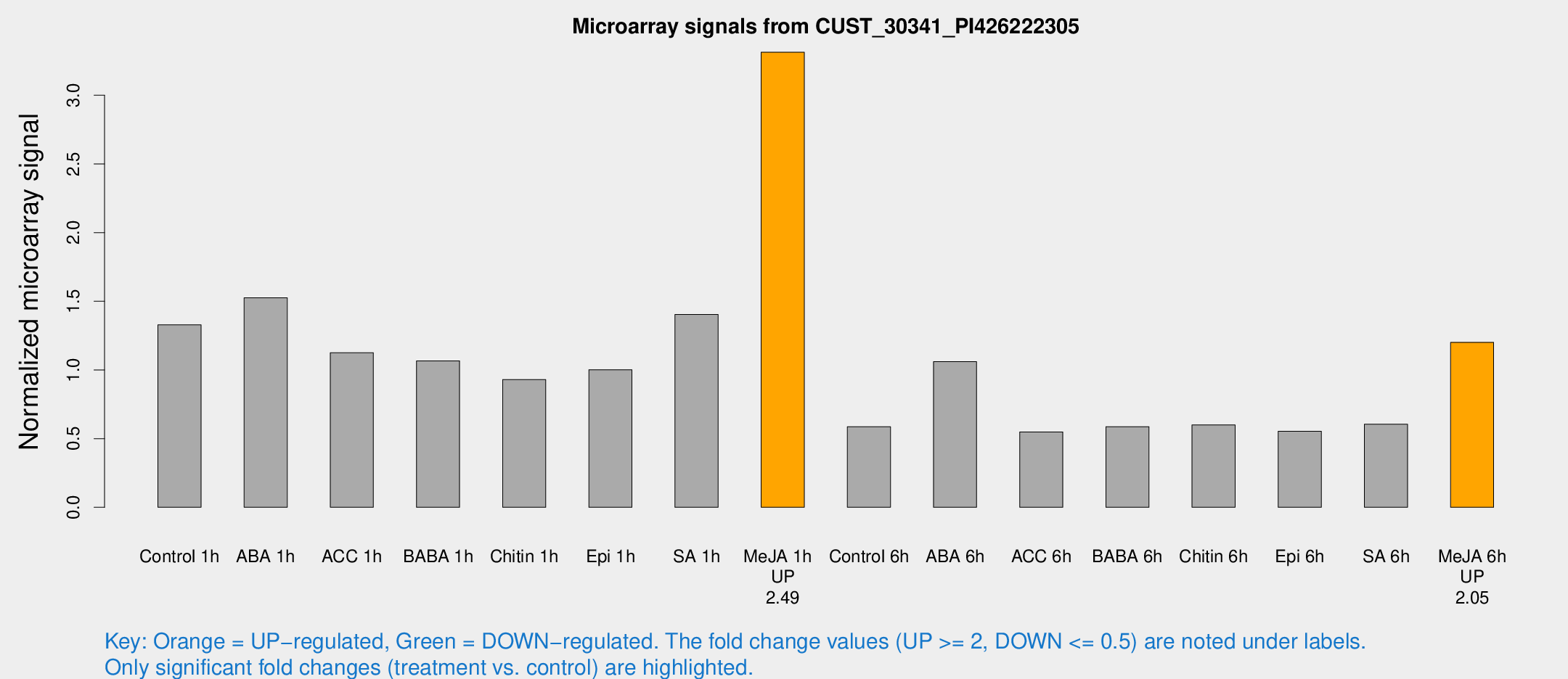
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2608.36 | 481.344 | 1.32834 | 0.155151 |
| ABA 1h | 2632.29 | 412.942 | 1.52491 | 0.161461 |
| ACC 1h | 2349.02 | 587.192 | 1.12503 | 0.234972 |
| BABA 1h | 2006.72 | 327.111 | 1.06625 | 0.0888929 |
| Chitin 1h | 1595.39 | 103.565 | 0.930169 | 0.0537352 |
| Epi 1h | 1652.12 | 95.7507 | 1.00173 | 0.057868 |
| SA 1h | 2860.13 | 617.953 | 1.40449 | 0.291134 |
| Me-JA 1h | 5196.24 | 559.82 | 3.3134 | 0.191311 |
| Control 6h | 1151.7 | 192.704 | 0.586933 | 0.0499108 |
| ABA 6h | 2149.73 | 135.959 | 1.06059 | 0.0612567 |
| ACC 6h | 1198.51 | 69.4863 | 0.548714 | 0.0664189 |
| BABA 6h | 1249.13 | 72.2581 | 0.587242 | 0.0339487 |
| Chitin 6h | 1215.62 | 70.4287 | 0.600073 | 0.0346915 |
| Epi 6h | 1187.92 | 68.8619 | 0.553409 | 0.0536094 |
| SA 6h | 1164.92 | 180.17 | 0.605055 | 0.0432229 |
| Me-JA 6h | 2271.3 | 131.188 | 1.20093 | 0.0963801 |
Source Transcript PGSC0003DMT400069793 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G55550.1 | +3 | 3e-171 | 508 | 252/399 (63%) | Concanavalin A-like lectin protein kinase family protein | chr3:20600019-20602073 REVERSE LENGTH=684 |
