Probe CUST_29790_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29790_PI426222305 | JHI_St_60k_v1 | DMT400051097 | TCTCCCGTGTAATTTGGCCTAGATCATGCCTCATGACAATGGCTTTCTCAAAACAGTAAG |
All Microarray Probes Designed to Gene DMG400019844
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29792_PI426222305 | JHI_St_60k_v1 | DMT400051098 | TCTCCCGTGTAATTTGGCCTAGATCATGCCTCATGACAATGGCTTTCTCAAAACAGTAAG |
| CUST_29790_PI426222305 | JHI_St_60k_v1 | DMT400051097 | TCTCCCGTGTAATTTGGCCTAGATCATGCCTCATGACAATGGCTTTCTCAAAACAGTAAG |
Microarray Signals from CUST_29790_PI426222305
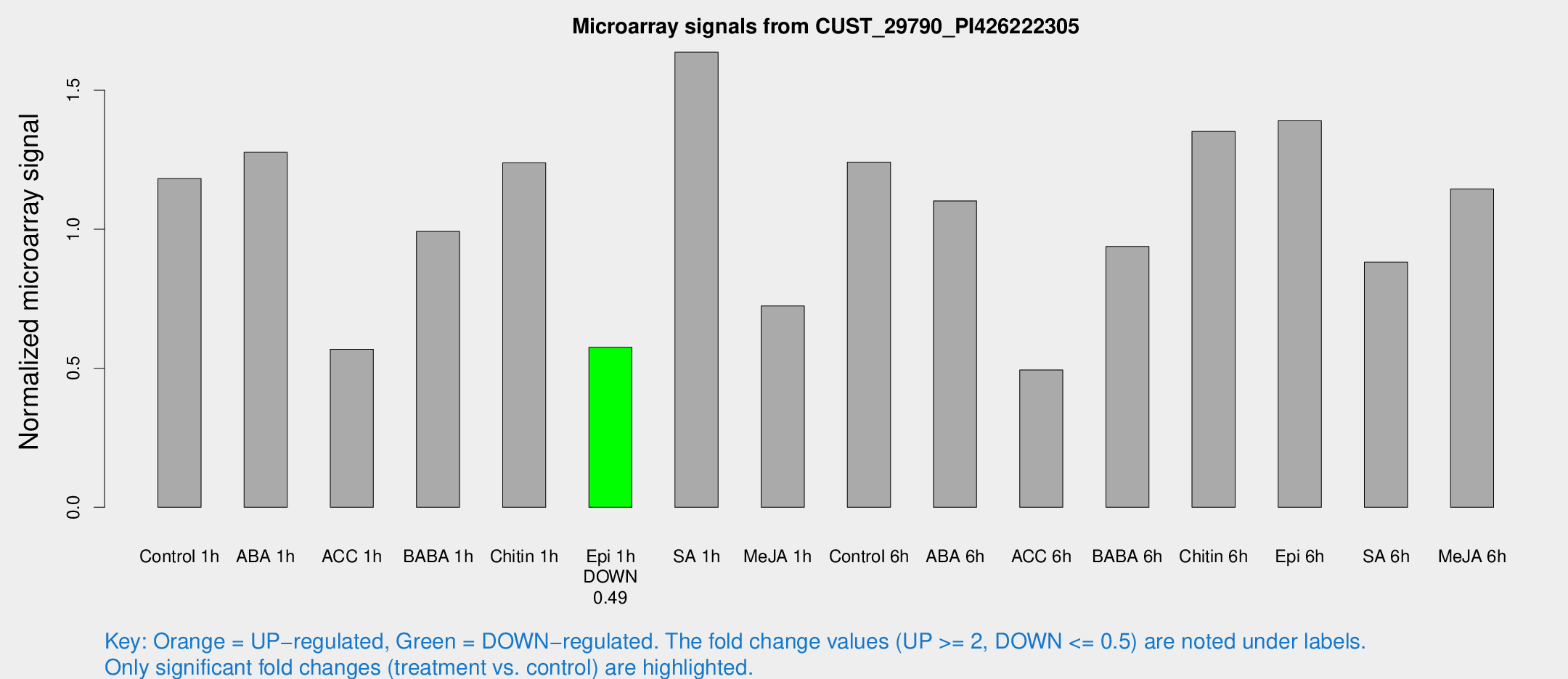
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 15.9479 | 3.15518 | 1.18214 | 0.240772 |
| ABA 1h | 16.4435 | 4.50723 | 1.27667 | 0.537097 |
| ACC 1h | 8.17793 | 3.48123 | 0.568475 | 0.270853 |
| BABA 1h | 15.193 | 5.72557 | 0.992289 | 0.383746 |
| Chitin 1h | 15.4646 | 3.26522 | 1.23872 | 0.344141 |
| Epi 1h | 6.70828 | 2.98139 | 0.575161 | 0.264187 |
| SA 1h | 22.3104 | 3.27433 | 1.63649 | 0.240188 |
| Me-JA 1h | 8.03478 | 3.09028 | 0.724169 | 0.300032 |
| Control 6h | 21.0326 | 7.92631 | 1.24102 | 0.680562 |
| ABA 6h | 16.7256 | 4.76967 | 1.10189 | 0.253032 |
| ACC 6h | 7.75886 | 3.70848 | 0.493818 | 0.244265 |
| BABA 6h | 16.6488 | 5.95293 | 0.937935 | 0.457587 |
| Chitin 6h | 20.2066 | 4.5451 | 1.35173 | 0.365043 |
| Epi 6h | 22.4426 | 6.49908 | 1.39029 | 0.412634 |
| SA 6h | 13.1898 | 4.34532 | 0.881983 | 0.502755 |
| Me-JA 6h | 17.8177 | 5.82203 | 1.14448 | 0.471984 |
Source Transcript PGSC0003DMT400051097 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G25770.3 | +3 | 1e-16 | 61 | 38/104 (37%) | alpha/beta-Hydrolases superfamily protein | chr5:8969308-8971806 REVERSE LENGTH=421 |
