Probe CUST_29778_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29778_PI426222305 | JHI_St_60k_v1 | DMT400053796 | GACAAACTTCAAAGTGTGTATCCTTATCTCATCAATGAAAGGCCAGCTTTTGTTTTAAAG |
All Microarray Probes Designed to Gene DMG400020870
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29778_PI426222305 | JHI_St_60k_v1 | DMT400053796 | GACAAACTTCAAAGTGTGTATCCTTATCTCATCAATGAAAGGCCAGCTTTTGTTTTAAAG |
| CUST_29788_PI426222305 | JHI_St_60k_v1 | DMT400053795 | GACAAACTTCAAAGTGTGTATCCTTATCTCATCAATGAAAGGCCAGCTTTTGTTTTAAAG |
Microarray Signals from CUST_29778_PI426222305
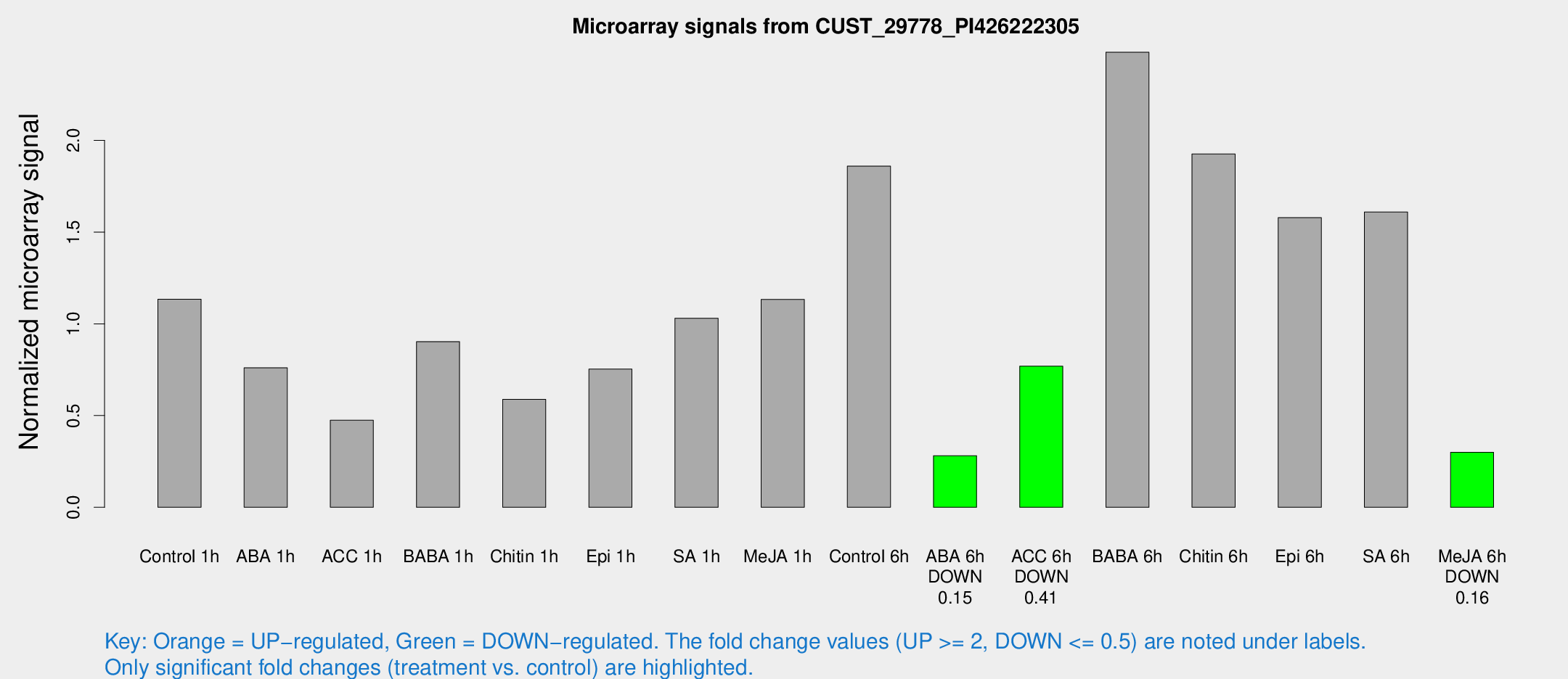
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 58.6943 | 13.7336 | 1.13363 | 0.184425 |
| ABA 1h | 35.0469 | 8.27594 | 0.760212 | 0.130437 |
| ACC 1h | 29.7641 | 10.7946 | 0.474339 | 0.243838 |
| BABA 1h | 43.2895 | 4.60259 | 0.902955 | 0.0974742 |
| Chitin 1h | 26.1798 | 3.77589 | 0.588472 | 0.0864127 |
| Epi 1h | 37.1142 | 11.8099 | 0.753226 | 0.335025 |
| SA 1h | 53.3391 | 7.65822 | 1.03062 | 0.213551 |
| Me-JA 1h | 57.0227 | 24.2523 | 1.13297 | 0.639844 |
| Control 6h | 95.405 | 17.2038 | 1.85951 | 0.20867 |
| ABA 6h | 15.0885 | 4.09137 | 0.281392 | 0.0816238 |
| ACC 6h | 47.7106 | 14.1472 | 0.76928 | 0.149959 |
| BABA 6h | 145.696 | 33.4346 | 2.48042 | 0.571732 |
| Chitin 6h | 101.329 | 7.23487 | 1.92559 | 0.17244 |
| Epi 6h | 89.684 | 13.7145 | 1.57879 | 0.29245 |
| SA 6h | 78.9339 | 6.5154 | 1.60912 | 0.327842 |
| Me-JA 6h | 18.6086 | 8.73458 | 0.299428 | 0.139915 |
Source Transcript PGSC0003DMT400053796 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G27910.1 | +3 | 7e-16 | 64 | 68/251 (27%) | SET domain protein 16 | chr4:13894694-13900256 FORWARD LENGTH=1027 |
