Probe CUST_29127_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29127_PI426222305 | JHI_St_60k_v1 | DMT400020647 | GTATTGGGCATCATCTATTGCTTCATAAAGTAATGAACTCCGGCGATTTTTGGTTGTTCA |
All Microarray Probes Designed to Gene DMG400008000
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29127_PI426222305 | JHI_St_60k_v1 | DMT400020647 | GTATTGGGCATCATCTATTGCTTCATAAAGTAATGAACTCCGGCGATTTTTGGTTGTTCA |
Microarray Signals from CUST_29127_PI426222305
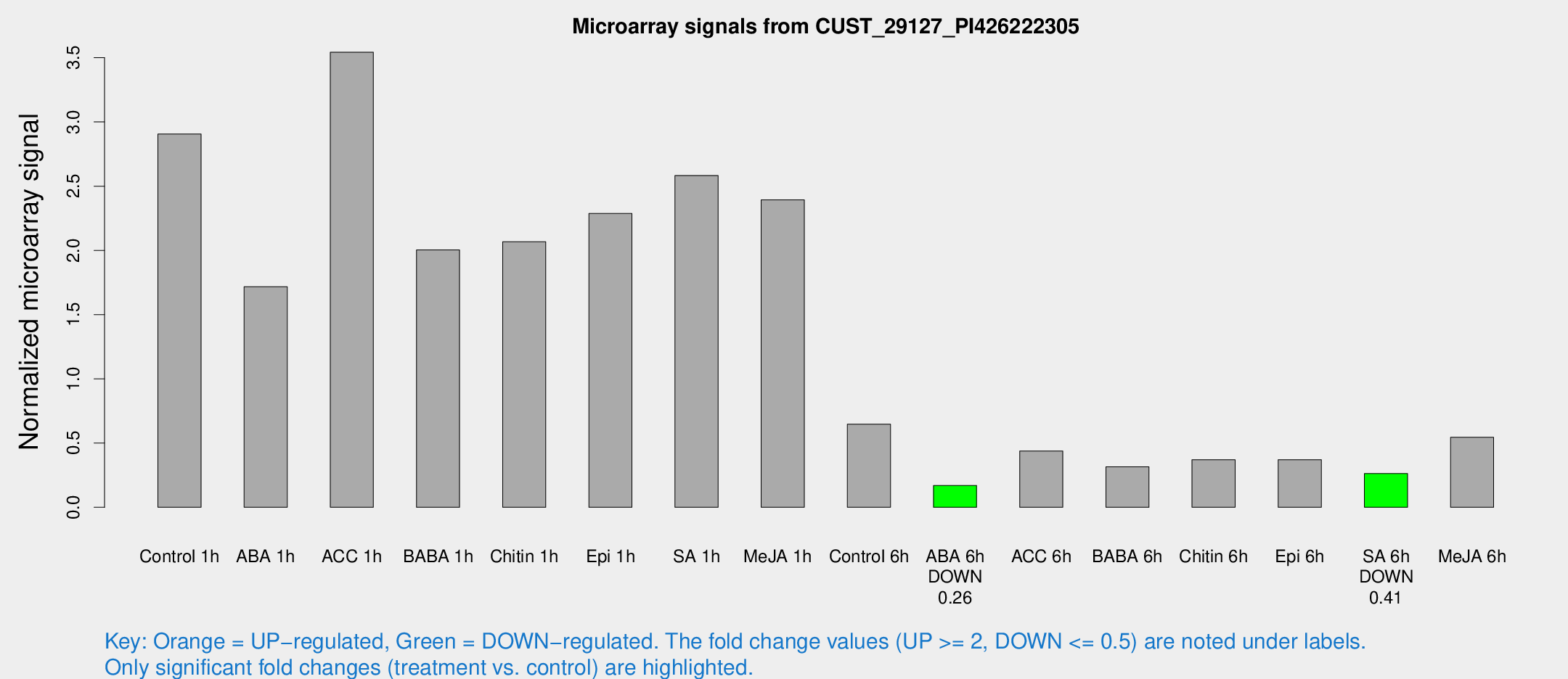
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11668.8 | 1977.21 | 2.90644 | 0.304193 |
| ABA 1h | 6043.42 | 807.665 | 1.71742 | 0.292901 |
| ACC 1h | 14952.2 | 3382.26 | 3.54263 | 0.577625 |
| BABA 1h | 7588.7 | 636.824 | 2.00346 | 0.115673 |
| Chitin 1h | 7729.51 | 1958.07 | 2.06686 | 0.657579 |
| Epi 1h | 7850.76 | 971.035 | 2.28719 | 0.278181 |
| SA 1h | 10545.9 | 1427.96 | 2.58295 | 0.475189 |
| Me-JA 1h | 7622 | 440.195 | 2.39304 | 0.228849 |
| Control 6h | 2776.27 | 729.934 | 0.647224 | 0.15499 |
| ABA 6h | 712.429 | 71.3738 | 0.17018 | 0.0101411 |
| ACC 6h | 2037.14 | 411.516 | 0.438505 | 0.0253319 |
| BABA 6h | 1399.85 | 169.928 | 0.315517 | 0.0303993 |
| Chitin 6h | 1565.82 | 200.8 | 0.370794 | 0.0372909 |
| Epi 6h | 1645.37 | 150.604 | 0.370793 | 0.0630382 |
| SA 6h | 1086.57 | 277.223 | 0.262455 | 0.0461537 |
| Me-JA 6h | 2167.71 | 313.373 | 0.54508 | 0.133775 |
Source Transcript PGSC0003DMT400020647 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G16150.1 | +1 | 0.0 | 547 | 270/325 (83%) | N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein | chr3:5471794-5473033 FORWARD LENGTH=325 |
