Probe CUST_29118_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29118_PI426222305 | JHI_St_60k_v1 | DMT400020515 | GGGTCTAATTACATAGGGTCCTTTTGGTGGTTGTTAATTAGCTTTAACTTGTTGATAAAG |
All Microarray Probes Designed to Gene DMG400007946
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28981_PI426222305 | JHI_St_60k_v1 | DMT400020516 | GGGTCTAATTACATAGGGTCCTTTTGGTGGTTGTTAATTAGCTTTAACTTGTTGATAAAG |
| CUST_29118_PI426222305 | JHI_St_60k_v1 | DMT400020515 | GGGTCTAATTACATAGGGTCCTTTTGGTGGTTGTTAATTAGCTTTAACTTGTTGATAAAG |
Microarray Signals from CUST_29118_PI426222305
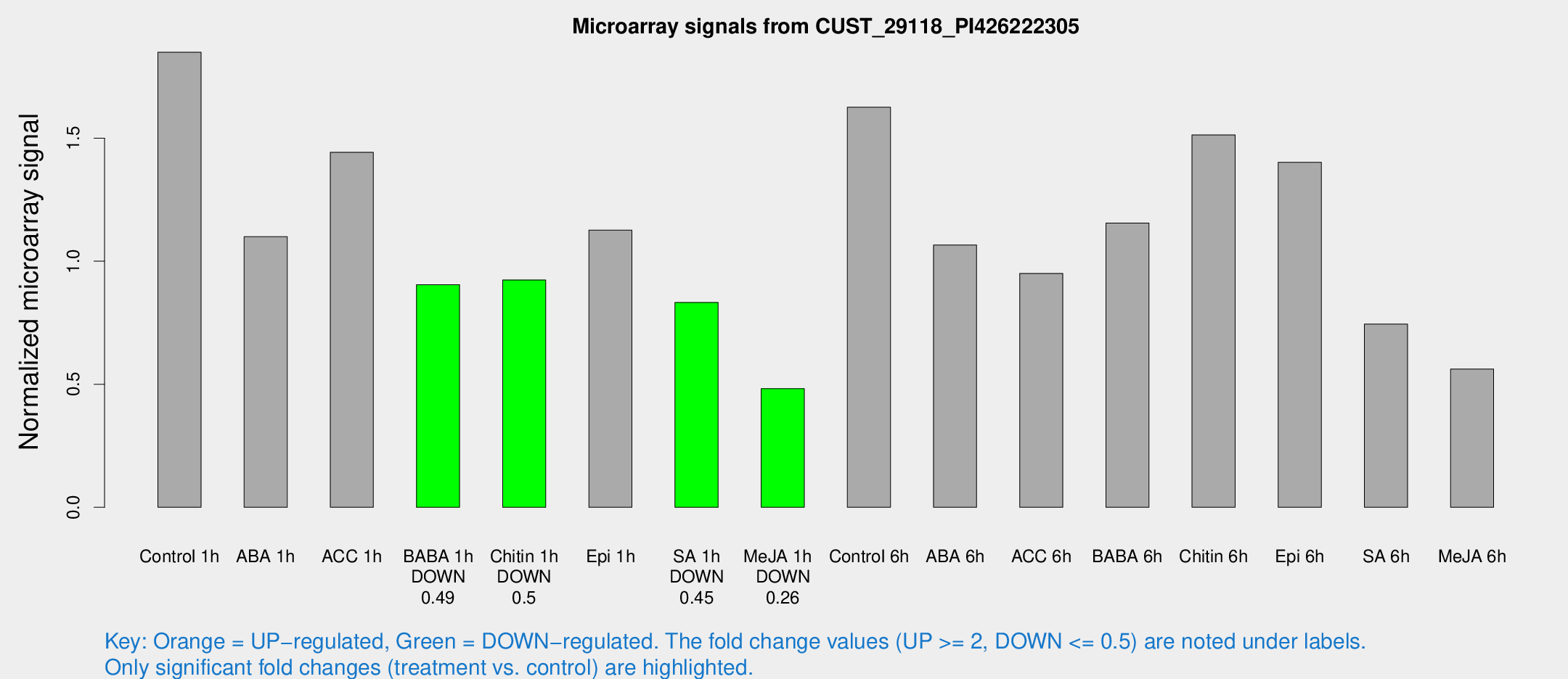
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1207.05 | 124.911 | 1.84902 | 0.106867 |
| ABA 1h | 651.077 | 114.013 | 1.09965 | 0.228154 |
| ACC 1h | 1084.88 | 351.446 | 1.44231 | 0.453897 |
| BABA 1h | 565.359 | 36.6454 | 0.904698 | 0.0640266 |
| Chitin 1h | 543.707 | 65.7608 | 0.92359 | 0.144163 |
| Epi 1h | 640.421 | 86.9402 | 1.12632 | 0.11942 |
| SA 1h | 552.651 | 32.0874 | 0.832475 | 0.058773 |
| Me-JA 1h | 259.244 | 37.5557 | 0.482313 | 0.108181 |
| Control 6h | 1133.67 | 273.921 | 1.6255 | 0.323133 |
| ABA 6h | 776.567 | 197.045 | 1.06589 | 0.317543 |
| ACC 6h | 735.999 | 163.047 | 0.950236 | 0.070109 |
| BABA 6h | 875.558 | 185.903 | 1.15512 | 0.230645 |
| Chitin 6h | 1062.67 | 157.521 | 1.513 | 0.252872 |
| Epi 6h | 1120.75 | 298.254 | 1.40143 | 0.625734 |
| SA 6h | 505.411 | 123.141 | 0.744715 | 0.121326 |
| Me-JA 6h | 410.44 | 141.713 | 0.561749 | 0.166273 |
Source Transcript PGSC0003DMT400020515 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G60060.1 | +2 | 4e-63 | 216 | 109/206 (53%) | Serine/threonine-protein kinase WNK (With No Lysine)-related | chr1:22139282-22141585 FORWARD LENGTH=386 |
