Probe CUST_28915_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28915_PI426222305 | JHI_St_60k_v1 | DMT400025448 | TATGCAATCTGGTGAAGCAGAGTTCCAGTTTGATCGTCGCCATGTCAAAGCTGTCATGTT |
All Microarray Probes Designed to Gene DMG400009823
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28915_PI426222305 | JHI_St_60k_v1 | DMT400025448 | TATGCAATCTGGTGAAGCAGAGTTCCAGTTTGATCGTCGCCATGTCAAAGCTGTCATGTT |
Microarray Signals from CUST_28915_PI426222305
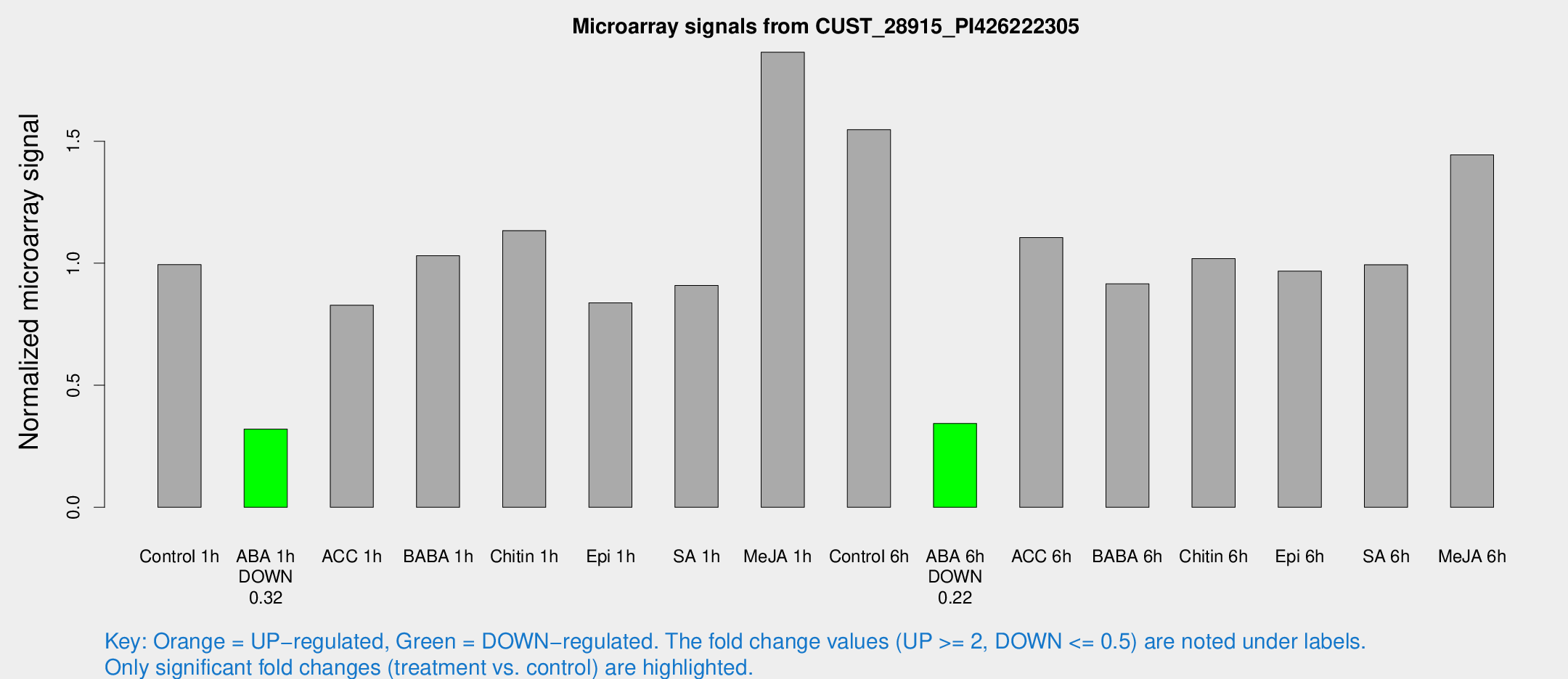
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 4325.19 | 633.808 | 0.994081 | 0.0772939 |
| ABA 1h | 1237.93 | 198.773 | 0.320616 | 0.0294814 |
| ACC 1h | 3830.49 | 809.984 | 0.828066 | 0.14218 |
| BABA 1h | 4530.91 | 1053.82 | 1.03057 | 0.177635 |
| Chitin 1h | 4362.81 | 282.78 | 1.13355 | 0.0712851 |
| Epi 1h | 3102.35 | 200.435 | 0.837529 | 0.0483642 |
| SA 1h | 4025.9 | 425.148 | 0.90866 | 0.0689447 |
| Me-JA 1h | 6770.26 | 1348.89 | 1.8645 | 0.234964 |
| Control 6h | 7259.31 | 2084.87 | 1.54741 | 0.385217 |
| ABA 6h | 1572.4 | 158.197 | 0.343394 | 0.0333745 |
| ACC 6h | 5608.48 | 1003.67 | 1.10545 | 0.148968 |
| BABA 6h | 4545.35 | 909.666 | 0.915609 | 0.166071 |
| Chitin 6h | 4655.52 | 395.89 | 1.0189 | 0.0588337 |
| Epi 6h | 4835.66 | 1001.94 | 0.967305 | 0.210351 |
| SA 6h | 4273.8 | 571.697 | 0.993764 | 0.0573837 |
| Me-JA 6h | 6227.39 | 801.279 | 1.444 | 0.169502 |
Source Transcript PGSC0003DMT400025448 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G48310.1 | +1 | 1e-42 | 152 | 80/228 (35%) | cytochrome P450, family 71, subfamily A, polypeptide 22 | chr3:17888192-17889749 FORWARD LENGTH=490 |
