Probe CUST_28740_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28740_PI426222305 | JHI_St_60k_v1 | DMT400010394 | GCTTCCATTTATTTCATCTGAAGTGTCTTGTAGAGTTGCTTTTTATATATGGCCTATCCA |
All Microarray Probes Designed to Gene DMG400004061
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28693_PI426222305 | JHI_St_60k_v1 | DMT400010395 | GCTTCCATTTATTTCATCTGAAGTGTCTTGTAGAGTTGCTTTTTATATATGGCCTATCCA |
| CUST_28740_PI426222305 | JHI_St_60k_v1 | DMT400010394 | GCTTCCATTTATTTCATCTGAAGTGTCTTGTAGAGTTGCTTTTTATATATGGCCTATCCA |
Microarray Signals from CUST_28740_PI426222305
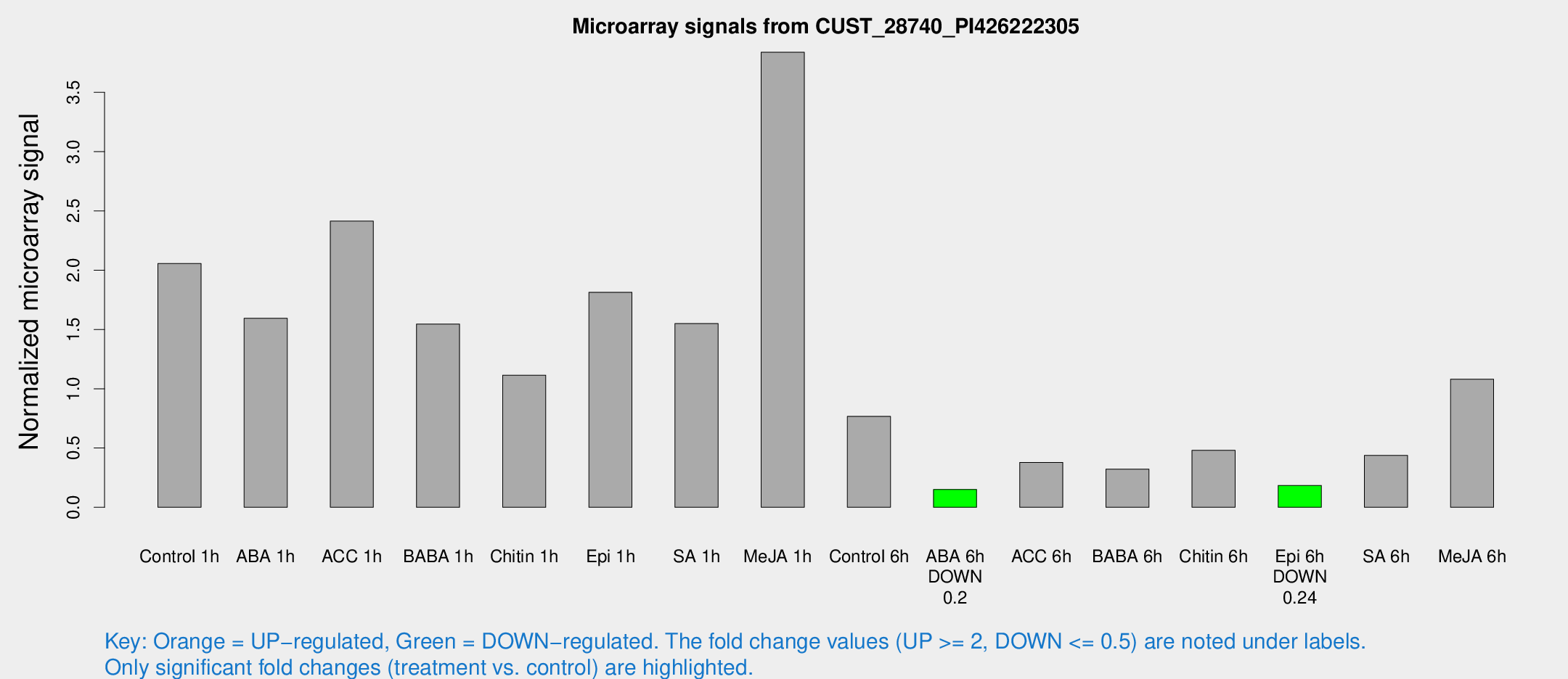
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 94.7251 | 25.2857 | 2.05697 | 0.45897 |
| ABA 1h | 62.9211 | 12.0332 | 1.59486 | 0.289102 |
| ACC 1h | 107.998 | 11.9771 | 2.41427 | 0.19488 |
| BABA 1h | 65.822 | 9.98417 | 1.546 | 0.130682 |
| Chitin 1h | 44.3376 | 7.06552 | 1.11476 | 0.114541 |
| Epi 1h | 68.7956 | 9.02692 | 1.81383 | 0.239311 |
| SA 1h | 73.0185 | 16.9106 | 1.54951 | 0.354808 |
| Me-JA 1h | 139.891 | 25.7214 | 3.83858 | 0.422585 |
| Control 6h | 34.2175 | 5.88477 | 0.766996 | 0.106362 |
| ABA 6h | 6.90146 | 4.00419 | 0.150647 | 0.0873094 |
| ACC 6h | 19.2129 | 4.67721 | 0.378656 | 0.119963 |
| BABA 6h | 16.5194 | 4.45329 | 0.322137 | 0.10455 |
| Chitin 6h | 22.9338 | 4.75343 | 0.480917 | 0.113038 |
| Epi 6h | 9.22591 | 4.42655 | 0.184779 | 0.0916512 |
| SA 6h | 18.9686 | 4.23233 | 0.43774 | 0.101816 |
| Me-JA 6h | 46.8807 | 5.20575 | 1.08111 | 0.209674 |
Source Transcript PGSC0003DMT400010394 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G08130.5 | +2 | 2e-56 | 204 | 153/310 (49%) | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | chr5:2606655-2609571 REVERSE LENGTH=532 |
