Probe CUST_28693_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28693_PI426222305 | JHI_St_60k_v1 | DMT400010395 | GCTTCCATTTATTTCATCTGAAGTGTCTTGTAGAGTTGCTTTTTATATATGGCCTATCCA |
All Microarray Probes Designed to Gene DMG400004061
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28693_PI426222305 | JHI_St_60k_v1 | DMT400010395 | GCTTCCATTTATTTCATCTGAAGTGTCTTGTAGAGTTGCTTTTTATATATGGCCTATCCA |
| CUST_28740_PI426222305 | JHI_St_60k_v1 | DMT400010394 | GCTTCCATTTATTTCATCTGAAGTGTCTTGTAGAGTTGCTTTTTATATATGGCCTATCCA |
Microarray Signals from CUST_28693_PI426222305
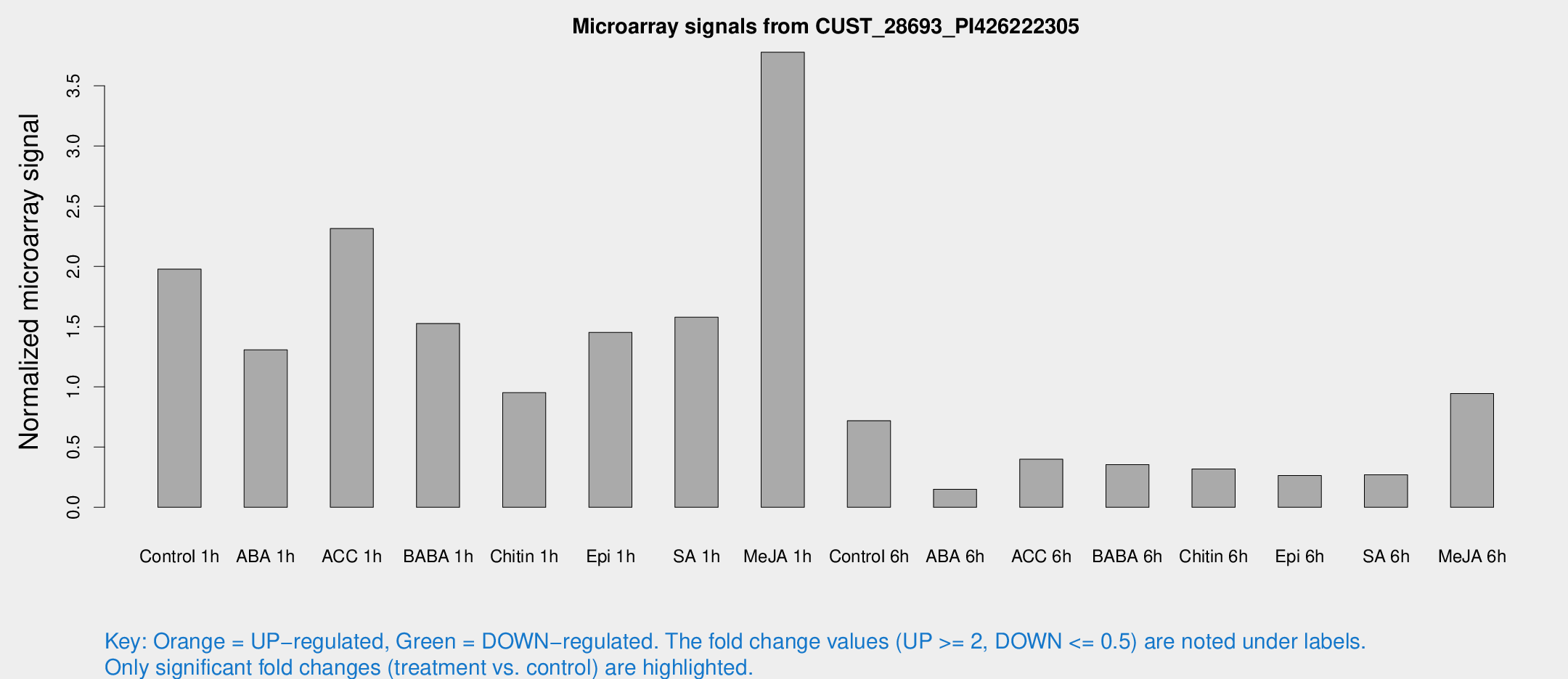
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 100.954 | 32.0573 | 1.97737 | 0.531036 |
| ABA 1h | 53.9234 | 4.3325 | 1.30826 | 0.105142 |
| ACC 1h | 113.347 | 18.8406 | 2.31466 | 0.375397 |
| BABA 1h | 69.3789 | 8.42031 | 1.52663 | 0.116949 |
| Chitin 1h | 40.6638 | 5.71115 | 0.951713 | 0.0927802 |
| Epi 1h | 58.9652 | 5.57251 | 1.45302 | 0.138026 |
| SA 1h | 81.5087 | 21.3448 | 1.57819 | 0.419189 |
| Me-JA 1h | 145.871 | 18.264 | 3.77917 | 0.240153 |
| Control 6h | 37.4976 | 12.3791 | 0.719186 | 0.207522 |
| ABA 6h | 7.86282 | 3.34673 | 0.150107 | 0.0725833 |
| ACC 6h | 22.1411 | 4.11582 | 0.399461 | 0.0800738 |
| BABA 6h | 19.2299 | 3.86906 | 0.355152 | 0.0769053 |
| Chitin 6h | 16.8952 | 4.14201 | 0.317952 | 0.0974791 |
| Epi 6h | 14.9789 | 3.89273 | 0.263565 | 0.101581 |
| SA 6h | 14.5528 | 5.5087 | 0.270696 | 0.0955149 |
| Me-JA 6h | 43.9102 | 4.09351 | 0.944143 | 0.0920917 |
Source Transcript PGSC0003DMT400010395 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G08130.5 | +3 | 4e-49 | 138 | 99/220 (45%) | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | chr5:2606655-2609571 REVERSE LENGTH=532 |
