Probe CUST_2845_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2845_PI426222305 | JHI_St_60k_v1 | DMT400000129 | CCGCGAAAAAAGGCATTGAAATTGTTACATTGATCGACCTACGATTGTTAAAGTTAAAAG |
All Microarray Probes Designed to Gene DMG400000037
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2845_PI426222305 | JHI_St_60k_v1 | DMT400000129 | CCGCGAAAAAAGGCATTGAAATTGTTACATTGATCGACCTACGATTGTTAAAGTTAAAAG |
Microarray Signals from CUST_2845_PI426222305
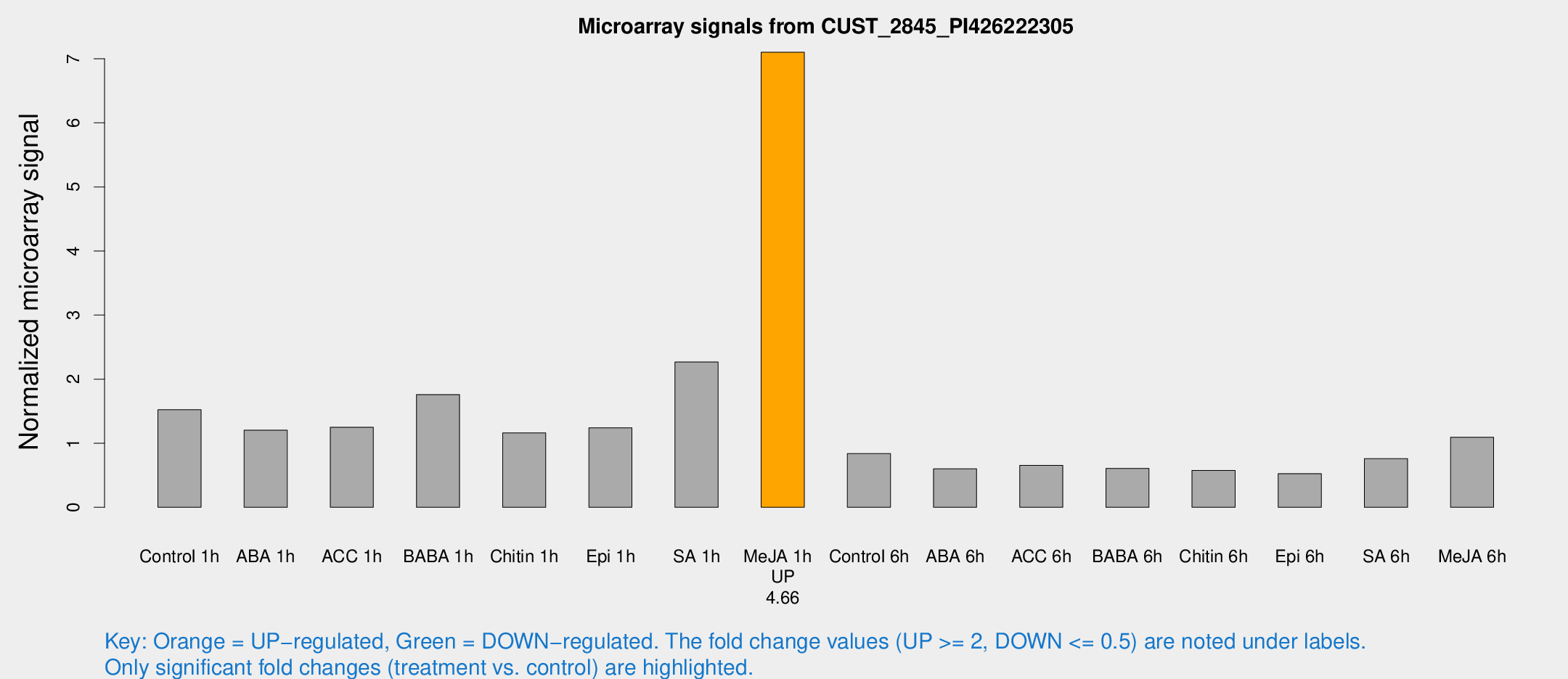
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2453.08 | 398.536 | 1.52267 | 0.175989 |
| ABA 1h | 1816.03 | 538.511 | 1.20369 | 0.264157 |
| ACC 1h | 2427.36 | 1053.23 | 1.25044 | 0.523379 |
| BABA 1h | 2706.55 | 319.128 | 1.75974 | 0.101626 |
| Chitin 1h | 1659.19 | 173.069 | 1.16051 | 0.0844271 |
| Epi 1h | 1717.94 | 215.07 | 1.24156 | 0.154586 |
| SA 1h | 4160.92 | 1553.14 | 2.26696 | 0.911473 |
| Me-JA 1h | 9286.15 | 1280.3 | 7.10203 | 0.410044 |
| Control 6h | 1374.6 | 265.775 | 0.837612 | 0.0963603 |
| ABA 6h | 1026.37 | 155.762 | 0.601302 | 0.0572839 |
| ACC 6h | 1192.75 | 111.644 | 0.655791 | 0.118435 |
| BABA 6h | 1072.96 | 76.4128 | 0.606801 | 0.0450556 |
| Chitin 6h | 986.792 | 155.902 | 0.576059 | 0.0967292 |
| Epi 6h | 948.633 | 135.638 | 0.525254 | 0.0819667 |
| SA 6h | 1241.72 | 284.245 | 0.759206 | 0.112464 |
| Me-JA 6h | 1718.12 | 99.5662 | 1.09429 | 0.157222 |
Source Transcript PGSC0003DMT400000129 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G01710.1 | +1 | 3e-49 | 170 | 77/107 (72%) | Acyl-CoA thioesterase family protein | chr1:262950-266029 FORWARD LENGTH=427 |
