Probe CUST_2805_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2805_PI426222305 | JHI_St_60k_v1 | DMT400000266 | GATTGAGGAAATACTATGGAACCGAAATGTTTTGGTTTGGACGATGAAAGTGATTTGATA |
All Microarray Probes Designed to Gene DMG400000088
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3121_PI426222305 | JHI_St_60k_v1 | DMT400000268 | GATTGAGGAAATACTATGGAACCGAAATGTTTTGGTTTGGACGATGAAAGTGATTTGATA |
| CUST_2805_PI426222305 | JHI_St_60k_v1 | DMT400000266 | GATTGAGGAAATACTATGGAACCGAAATGTTTTGGTTTGGACGATGAAAGTGATTTGATA |
Microarray Signals from CUST_2805_PI426222305
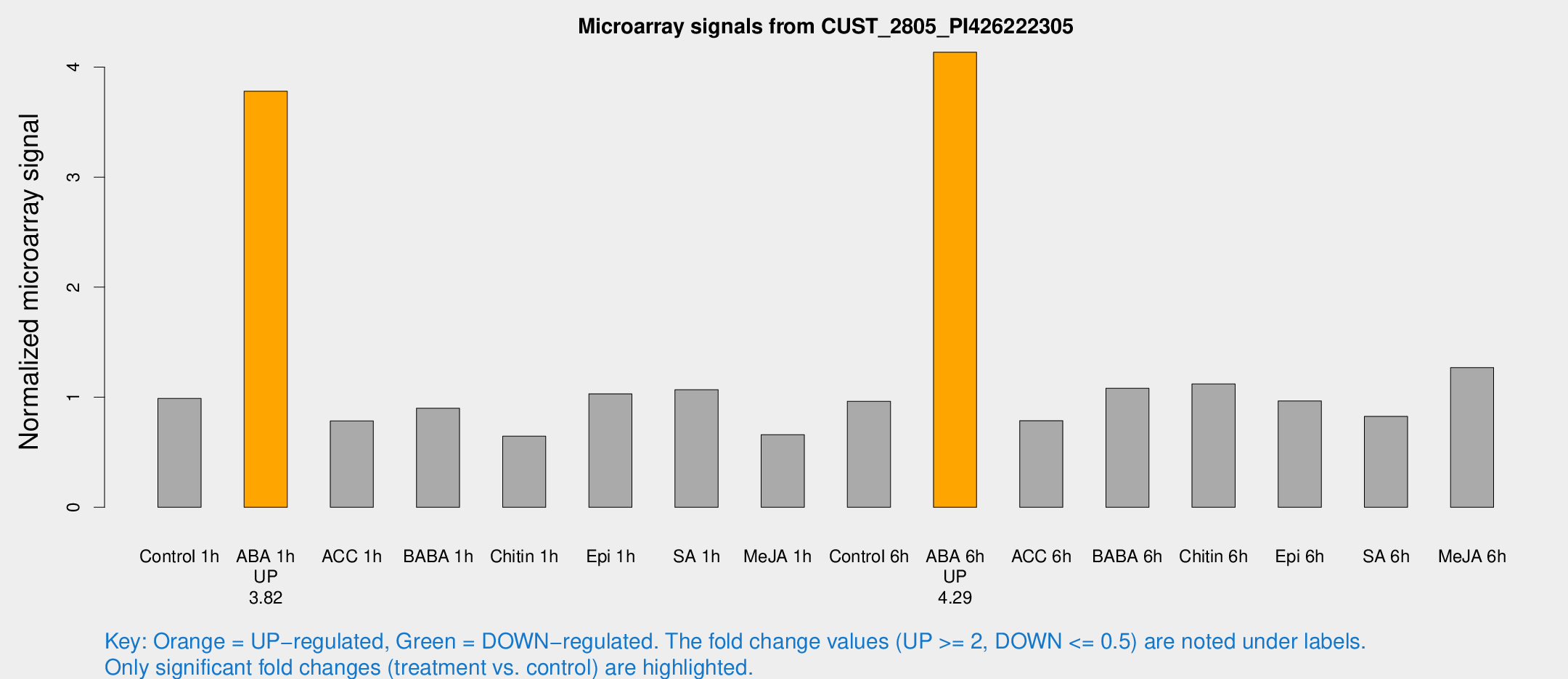
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 840.474 | 115.262 | 0.988493 | 0.0874415 |
| ABA 1h | 2842.73 | 358.638 | 3.77965 | 0.350247 |
| ACC 1h | 780.26 | 249.793 | 0.784191 | 0.293527 |
| BABA 1h | 759.904 | 150.33 | 0.899929 | 0.110288 |
| Chitin 1h | 497.048 | 79.1336 | 0.645955 | 0.0734296 |
| Epi 1h | 766.43 | 122.811 | 1.03048 | 0.158649 |
| SA 1h | 946.139 | 169.5 | 1.06886 | 0.122772 |
| Me-JA 1h | 459.304 | 62.2182 | 0.65985 | 0.121488 |
| Control 6h | 848.82 | 180.303 | 0.963578 | 0.149061 |
| ABA 6h | 3716.94 | 400.382 | 4.13454 | 0.243976 |
| ACC 6h | 767.593 | 97.6709 | 0.78683 | 0.0782874 |
| BABA 6h | 1035.65 | 162.985 | 1.08157 | 0.149985 |
| Chitin 6h | 1002.25 | 75.6366 | 1.12093 | 0.114057 |
| Epi 6h | 916.856 | 74.8433 | 0.966576 | 0.145516 |
| SA 6h | 771.059 | 227.887 | 0.825572 | 0.225001 |
| Me-JA 6h | 1147.46 | 331.31 | 1.2692 | 0.268418 |
Source Transcript PGSC0003DMT400000266 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G46270.1 | +1 | 4e-102 | 318 | 220/425 (52%) | G-box binding factor 3 | chr2:19000859-19002901 FORWARD LENGTH=382 |
