Probe CUST_27887_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27887_PI426222305 | JHI_St_60k_v1 | DMT400081935 | GGATTTAGTACTACAAATGTTGCTCTTAAGCAAGGCATTACCATGTTCCGTTCTTAAATA |
All Microarray Probes Designed to Gene DMG400032163
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27976_PI426222305 | JHI_St_60k_v1 | DMT400081936 | GGATTTAGTACTACAAATGTTGCTCTTAAGCAAGGCATTACCATGTTCCGTTCTTAAATA |
| CUST_27887_PI426222305 | JHI_St_60k_v1 | DMT400081935 | GGATTTAGTACTACAAATGTTGCTCTTAAGCAAGGCATTACCATGTTCCGTTCTTAAATA |
Microarray Signals from CUST_27887_PI426222305
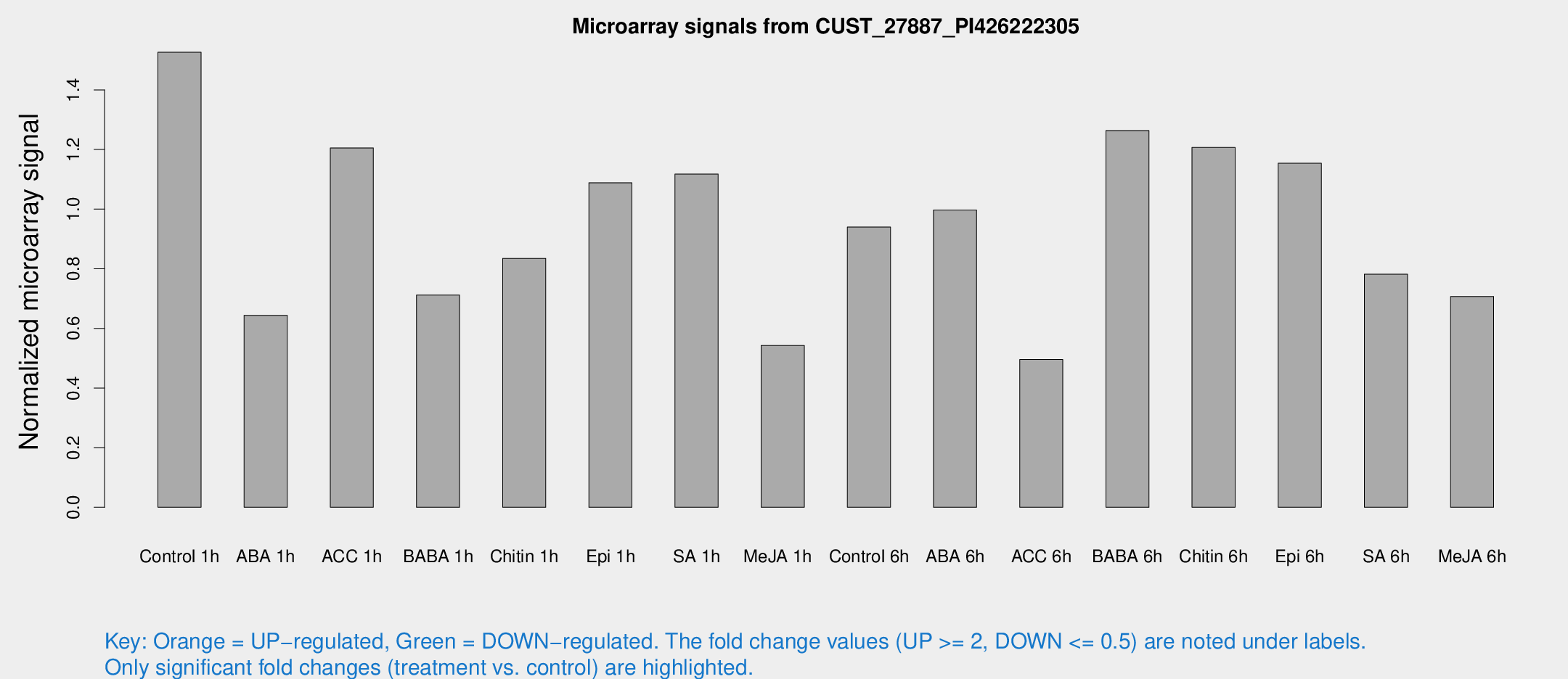
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 26.5202 | 5.68489 | 1.52631 | 0.221097 |
| ABA 1h | 10.7624 | 3.73605 | 0.644037 | 0.243287 |
| ACC 1h | 27.5986 | 14.4057 | 1.20536 | 0.69124 |
| BABA 1h | 11.7368 | 3.42602 | 0.71197 | 0.226938 |
| Chitin 1h | 12.7733 | 3.25173 | 0.834893 | 0.227271 |
| Epi 1h | 15.839 | 3.24968 | 1.08844 | 0.230169 |
| SA 1h | 21.1526 | 5.82757 | 1.11784 | 0.415168 |
| Me-JA 1h | 7.58506 | 3.31124 | 0.542588 | 0.252794 |
| Control 6h | 19.3266 | 6.92133 | 0.940126 | 0.448855 |
| ABA 6h | 19.7056 | 6.10836 | 0.997347 | 0.405056 |
| ACC 6h | 12.5203 | 6.82631 | 0.495716 | 0.232179 |
| BABA 6h | 36.2762 | 19.2563 | 1.26361 | 1.32467 |
| Chitin 6h | 25.3602 | 9.82464 | 1.20701 | 0.566131 |
| Epi 6h | 24.7399 | 7.77254 | 1.15387 | 0.602215 |
| SA 6h | 14.498 | 4.32501 | 0.78204 | 0.268348 |
| Me-JA 6h | 13.5749 | 4.53726 | 0.706969 | 0.249377 |
Source Transcript PGSC0003DMT400081935 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G51680.1 | +1 | 3e-69 | 220 | 131/266 (49%) | NAD(P)-binding Rossmann-fold superfamily protein | chr3:19173622-19174667 REVERSE LENGTH=303 |
