Probe CUST_27844_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27844_PI426222305 | JHI_St_60k_v1 | DMT400015940 | TCACAGAGCATTTGAGAGAAGAAGAAGGAAGAAGATTGTATTAATGTCTTGATTCAGTGA |
All Microarray Probes Designed to Gene DMG400006237
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27841_PI426222305 | JHI_St_60k_v1 | DMT400015939 | AAAGCTAAAAGTTCCAACAATGCCAAAGCCTGAAATCCCAGAACTGCCTAAACCAACTTT |
| CUST_27844_PI426222305 | JHI_St_60k_v1 | DMT400015940 | TCACAGAGCATTTGAGAGAAGAAGAAGGAAGAAGATTGTATTAATGTCTTGATTCAGTGA |
Microarray Signals from CUST_27844_PI426222305
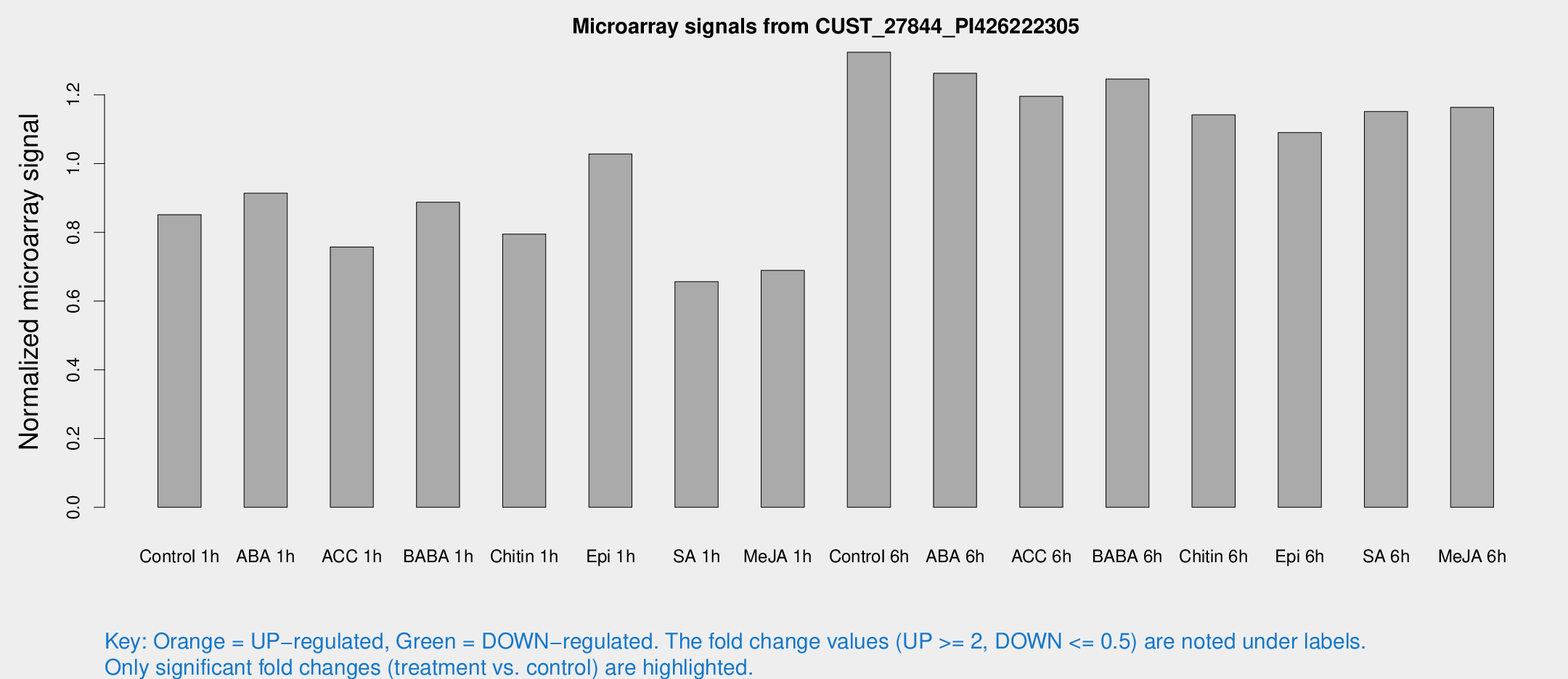
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 44.8781 | 9.993 | 0.851235 | 0.168944 |
| ABA 1h | 43.5375 | 12.9281 | 0.913972 | 0.268598 |
| ACC 1h | 42.4038 | 11.5529 | 0.757629 | 0.196254 |
| BABA 1h | 42.8219 | 4.40613 | 0.887503 | 0.091296 |
| Chitin 1h | 36.2227 | 5.26977 | 0.794949 | 0.0908532 |
| Epi 1h | 45.7669 | 8.88969 | 1.0282 | 0.202148 |
| SA 1h | 37.2386 | 11.7472 | 0.656643 | 0.185192 |
| Me-JA 1h | 28.6484 | 4.37816 | 0.689311 | 0.14881 |
| Control 6h | 70.1627 | 15.6688 | 1.32411 | 0.224489 |
| ABA 6h | 70.9674 | 17.733 | 1.26275 | 0.286762 |
| ACC 6h | 69.1404 | 8.25951 | 1.19611 | 0.100827 |
| BABA 6h | 72.4156 | 14.4599 | 1.24626 | 0.221743 |
| Chitin 6h | 61.894 | 9.10021 | 1.14201 | 0.192922 |
| Epi 6h | 61.8352 | 6.82962 | 1.0903 | 0.122062 |
| SA 6h | 56.9786 | 5.08726 | 1.15138 | 0.149439 |
| Me-JA 6h | 61.3217 | 14.7587 | 1.16371 | 0.20347 |
Source Transcript PGSC0003DMT400015940 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G09530.1 | +3 | 2e-39 | 107 | 64/163 (39%) | hydroxyproline-rich glycoprotein family protein | chr5:2960080-2961192 REVERSE LENGTH=370 |
