Probe CUST_27483_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27483_PI426222305 | JHI_St_60k_v1 | DMT400024494 | GCGATTTACTGGTGTTCTATTCCTTGAAAACCACCTGCTATGATTCTTGAATCTAATTAT |
All Microarray Probes Designed to Gene DMG400009472
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27419_PI426222305 | JHI_St_60k_v1 | DMT400024493 | CTTTGGTTTCTTCACCAACTCACAAGCATCTTCTCGCATCGTTTGGCTGTATAAATAAGC |
| CUST_27483_PI426222305 | JHI_St_60k_v1 | DMT400024494 | GCGATTTACTGGTGTTCTATTCCTTGAAAACCACCTGCTATGATTCTTGAATCTAATTAT |
| CUST_27459_PI426222305 | JHI_St_60k_v1 | DMT400024492 | CTTTGGTTTCTTCACCAACTCACAAGCATCTTCTCGCATCGTTTGGCTGTATAAATAAGC |
Microarray Signals from CUST_27483_PI426222305
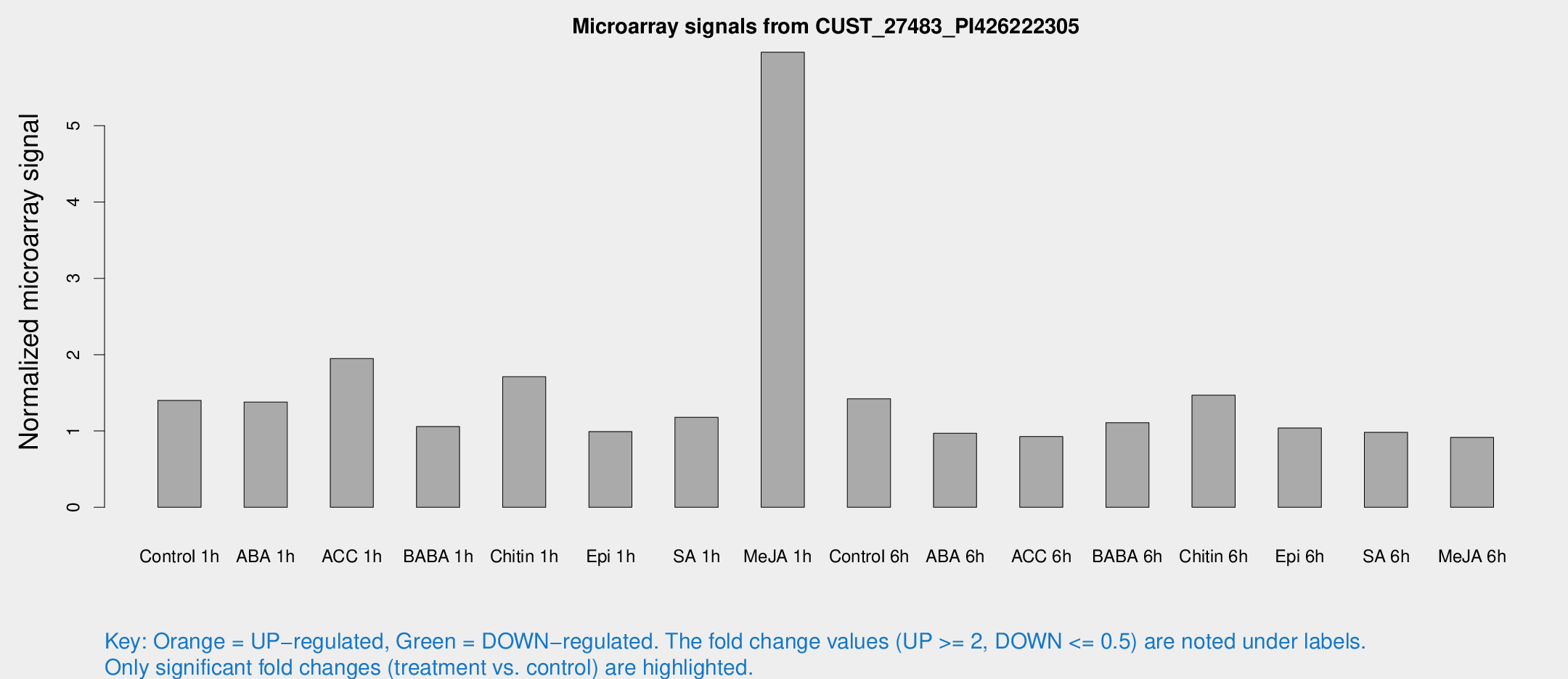
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 12.8943 | 6.69717 | 1.40006 | 0.682214 |
| ABA 1h | 10.2025 | 4.21765 | 1.37888 | 0.645588 |
| ACC 1h | 16.8323 | 6.77439 | 1.94936 | 0.704629 |
| BABA 1h | 7.35659 | 3.91981 | 1.0582 | 0.571065 |
| Chitin 1h | 11.8125 | 3.7643 | 1.71063 | 0.648443 |
| Epi 1h | 6.15622 | 3.57242 | 0.991781 | 0.574873 |
| SA 1h | 9.68294 | 3.7257 | 1.18013 | 0.55982 |
| Me-JA 1h | 34.9544 | 4.29561 | 5.96282 | 0.730203 |
| Control 6h | 10.7302 | 3.90446 | 1.4228 | 0.580224 |
| ABA 6h | 7.49481 | 4.00427 | 0.970504 | 0.526416 |
| ACC 6h | 7.78275 | 4.61222 | 0.927052 | 0.537089 |
| BABA 6h | 9.02275 | 4.39374 | 1.10882 | 0.561071 |
| Chitin 6h | 12.0556 | 4.35112 | 1.46915 | 0.624525 |
| Epi 6h | 8.50284 | 4.46101 | 1.03874 | 0.552206 |
| SA 6h | 6.96005 | 4.03282 | 0.981723 | 0.568497 |
| Me-JA 6h | 6.53725 | 3.79116 | 0.915744 | 0.530661 |
Source Transcript PGSC0003DMT400024494 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
