Probe CUST_27128_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27128_PI426222305 | JHI_St_60k_v1 | DMT400026311 | GTACAACAGTTACCTTGGTTGTATCTGTAAAGTTTATCACTTTCAGTATTTCCAGTGAGA |
All Microarray Probes Designed to Gene DMG400010154
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27128_PI426222305 | JHI_St_60k_v1 | DMT400026311 | GTACAACAGTTACCTTGGTTGTATCTGTAAAGTTTATCACTTTCAGTATTTCCAGTGAGA |
| CUST_27105_PI426222305 | JHI_St_60k_v1 | DMT400026310 | TTATTAAGAAGATCAACCATGAGAGATTGCTGCAGAGGCAAGAATCCGATCAGCGTGCCT |
Microarray Signals from CUST_27128_PI426222305
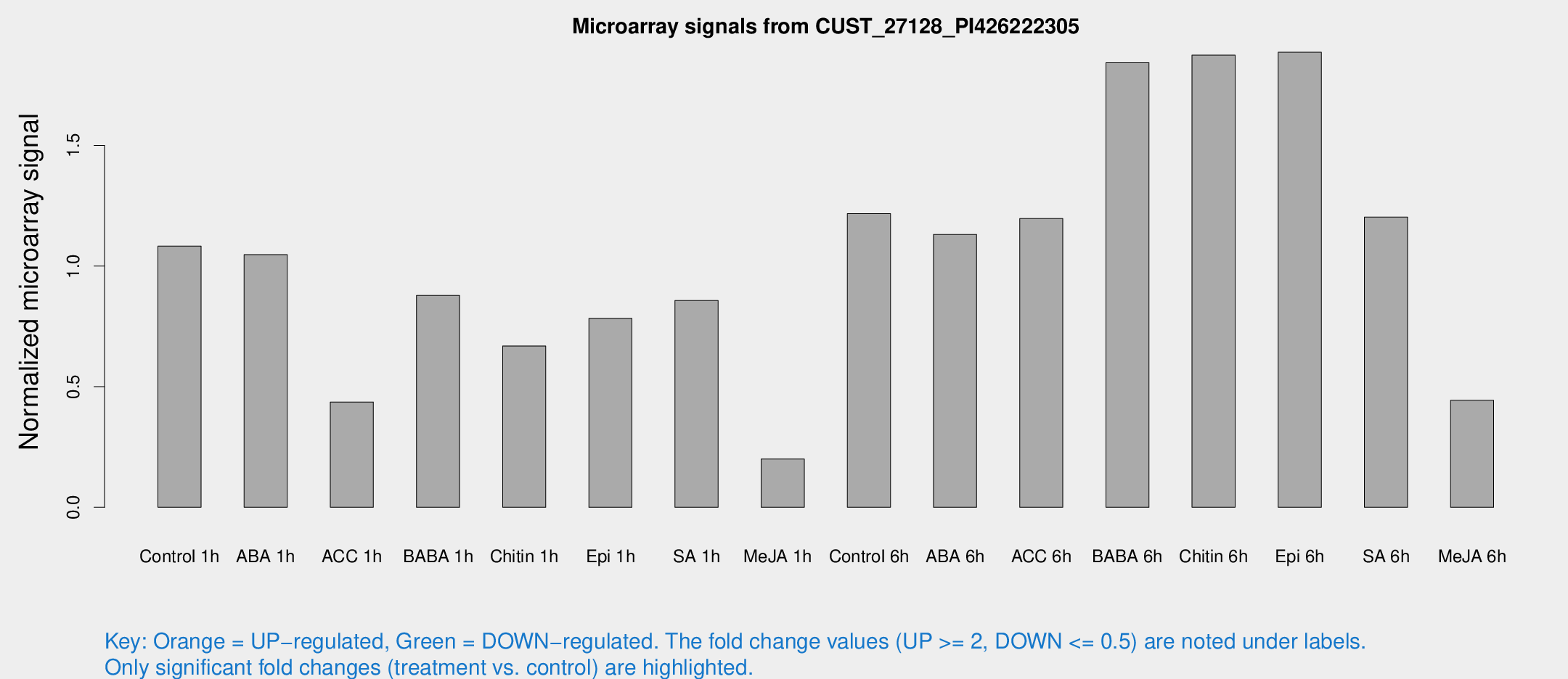
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 63.9982 | 15.2229 | 1.08332 | 0.184627 |
| ABA 1h | 56.656 | 16.4914 | 1.04754 | 0.255032 |
| ACC 1h | 34.8481 | 14.1605 | 0.436584 | 0.342665 |
| BABA 1h | 51.6569 | 13.4116 | 0.878237 | 0.189357 |
| Chitin 1h | 35.4233 | 7.50863 | 0.6688 | 0.160227 |
| Epi 1h | 38.299 | 3.94666 | 0.783293 | 0.0817895 |
| SA 1h | 51.1438 | 9.23474 | 0.857444 | 0.179792 |
| Me-JA 1h | 10.747 | 4.49561 | 0.200255 | 0.08442 |
| Control 6h | 76.5048 | 21.8255 | 1.21771 | 0.347913 |
| ABA 6h | 71.0944 | 15.3534 | 1.13136 | 0.309388 |
| ACC 6h | 81.5843 | 18.5522 | 1.19729 | 0.203153 |
| BABA 6h | 115.382 | 7.73548 | 1.84347 | 0.12355 |
| Chitin 6h | 111.634 | 7.49903 | 1.87481 | 0.125936 |
| Epi 6h | 119.524 | 8.53302 | 1.88683 | 0.186074 |
| SA 6h | 73.0465 | 19.9079 | 1.20302 | 0.255993 |
| Me-JA 6h | 28.4054 | 8.85262 | 0.443678 | 0.15653 |
Source Transcript PGSC0003DMT400026311 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
