Probe CUST_27067_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27067_PI426222305 | JHI_St_60k_v1 | DMT400065841 | GTTTAATGGATTTGATATGTTATCGACACCTACCTATCTTGATAGCACCAATTCTACCTA |
All Microarray Probes Designed to Gene DMG400025628
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27067_PI426222305 | JHI_St_60k_v1 | DMT400065841 | GTTTAATGGATTTGATATGTTATCGACACCTACCTATCTTGATAGCACCAATTCTACCTA |
Microarray Signals from CUST_27067_PI426222305
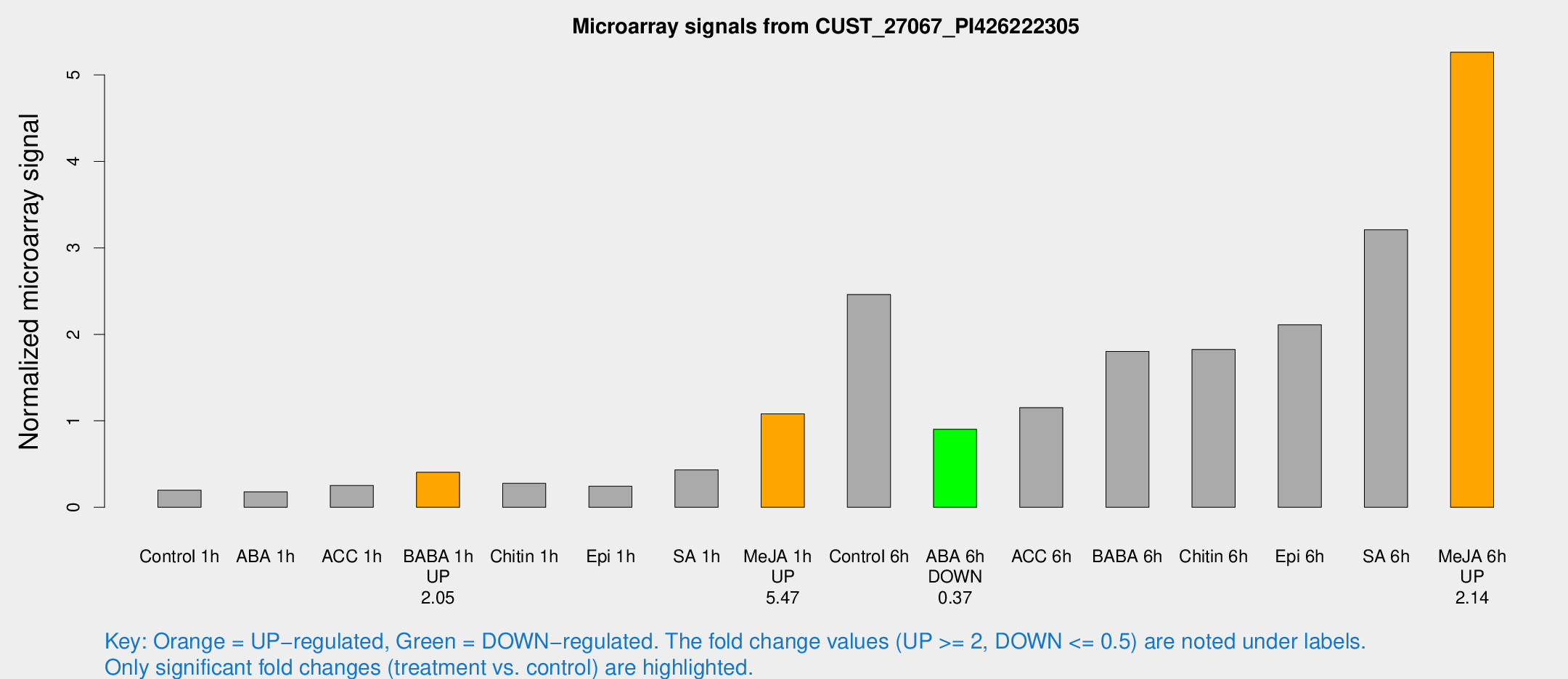
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 574.375 | 77.671 | 0.197408 | 0.013246 |
| ABA 1h | 517.599 | 195.798 | 0.178085 | 0.0570695 |
| ACC 1h | 767.206 | 151.714 | 0.252097 | 0.0424663 |
| BABA 1h | 1149.22 | 191.77 | 0.404566 | 0.0344911 |
| Chitin 1h | 720.06 | 77.521 | 0.277034 | 0.0348901 |
| Epi 1h | 637.695 | 135.312 | 0.244387 | 0.0556353 |
| SA 1h | 1374.82 | 385.853 | 0.432255 | 0.146696 |
| Me-JA 1h | 2546.49 | 259.525 | 1.0803 | 0.06372 |
| Control 6h | 7628.31 | 2159.54 | 2.46156 | 0.504044 |
| ABA 6h | 2780.94 | 329.019 | 0.902696 | 0.155629 |
| ACC 6h | 3812.38 | 347.101 | 1.15298 | 0.0665786 |
| BABA 6h | 5850.97 | 660.324 | 1.80285 | 0.152897 |
| Chitin 6h | 5773.11 | 1118.34 | 1.82584 | 0.406455 |
| Epi 6h | 7154.34 | 1468.36 | 2.11145 | 0.687345 |
| SA 6h | 9756.39 | 2342.35 | 3.20913 | 0.539002 |
| Me-JA 6h | 15870.8 | 3430.88 | 5.2624 | 0.979976 |
Source Transcript PGSC0003DMT400065841 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14390.1 | +3 | 3e-20 | 82 | 81/305 (27%) | Pyridoxal-dependent decarboxylase family protein | chr3:4806771-4808954 FORWARD LENGTH=484 |
