Probe CUST_26951_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26951_PI426222305 | JHI_St_60k_v1 | DMT400052686 | CAAGTACTCCAAAAGATTTGGATATGAGCGAGAAAAGTGGATTAAGTGCAGCGAAAGAGA |
All Microarray Probes Designed to Gene DMG402020453
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26951_PI426222305 | JHI_St_60k_v1 | DMT400052686 | CAAGTACTCCAAAAGATTTGGATATGAGCGAGAAAAGTGGATTAAGTGCAGCGAAAGAGA |
Microarray Signals from CUST_26951_PI426222305
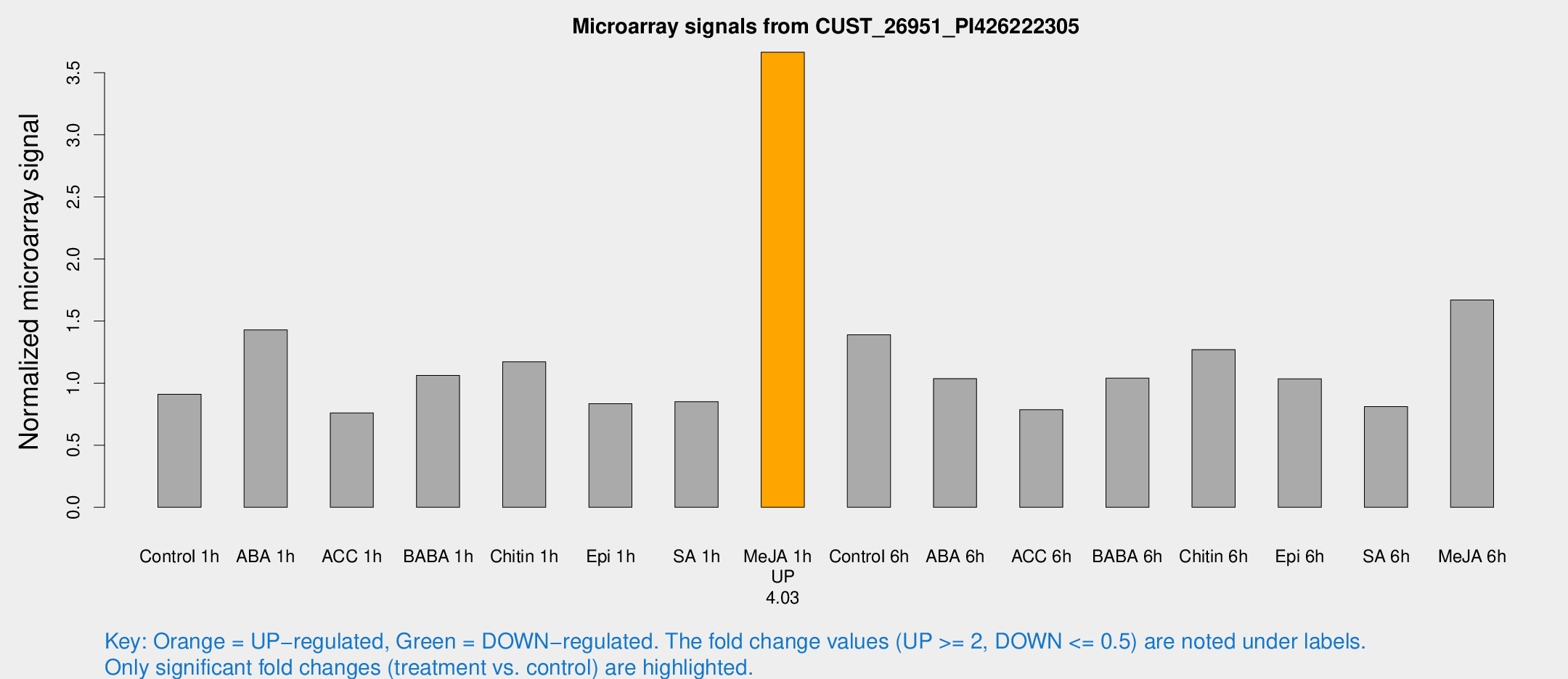
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 106.245 | 33.9345 | 0.910178 | 0.223124 |
| ABA 1h | 142.12 | 31.0057 | 1.4291 | 0.266155 |
| ACC 1h | 83.7583 | 7.951 | 0.760398 | 0.061456 |
| BABA 1h | 109.519 | 8.03105 | 1.06291 | 0.0711515 |
| Chitin 1h | 113.525 | 12.5446 | 1.17242 | 0.0763157 |
| Epi 1h | 78.0889 | 9.65515 | 0.834847 | 0.0938445 |
| SA 1h | 97.0199 | 20.5989 | 0.850817 | 0.206773 |
| Me-JA 1h | 323.508 | 41.4933 | 3.66511 | 0.215466 |
| Control 6h | 169.025 | 57.3463 | 1.38878 | 0.448374 |
| ABA 6h | 117.767 | 8.50443 | 1.03652 | 0.109545 |
| ACC 6h | 98.7133 | 17.4789 | 0.785853 | 0.056368 |
| BABA 6h | 129.127 | 27.7568 | 1.04026 | 0.214279 |
| Chitin 6h | 144.964 | 13.5367 | 1.26918 | 0.164951 |
| Epi 6h | 131.246 | 30.9957 | 1.03527 | 0.350729 |
| SA 6h | 91.0319 | 22.2958 | 0.810695 | 0.129715 |
| Me-JA 6h | 188.303 | 43.1694 | 1.66997 | 0.306726 |
Source Transcript PGSC0003DMT400052686 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G26330.1 | +1 | 3e-45 | 157 | 85/173 (49%) | cytochrome P450, family 71, subfamily B, polypeptide 37 | chr3:9646873-9648536 REVERSE LENGTH=500 |
