Probe CUST_26811_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26811_PI426222305 | JHI_St_60k_v1 | DMT400035281 | TAGCTCATGAGGGGAGGATAAGATATTAAGGAGTTAGTCCCTTCGATTATAAAGTTATAA |
All Microarray Probes Designed to Gene DMG400013557
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26811_PI426222305 | JHI_St_60k_v1 | DMT400035281 | TAGCTCATGAGGGGAGGATAAGATATTAAGGAGTTAGTCCCTTCGATTATAAAGTTATAA |
Microarray Signals from CUST_26811_PI426222305
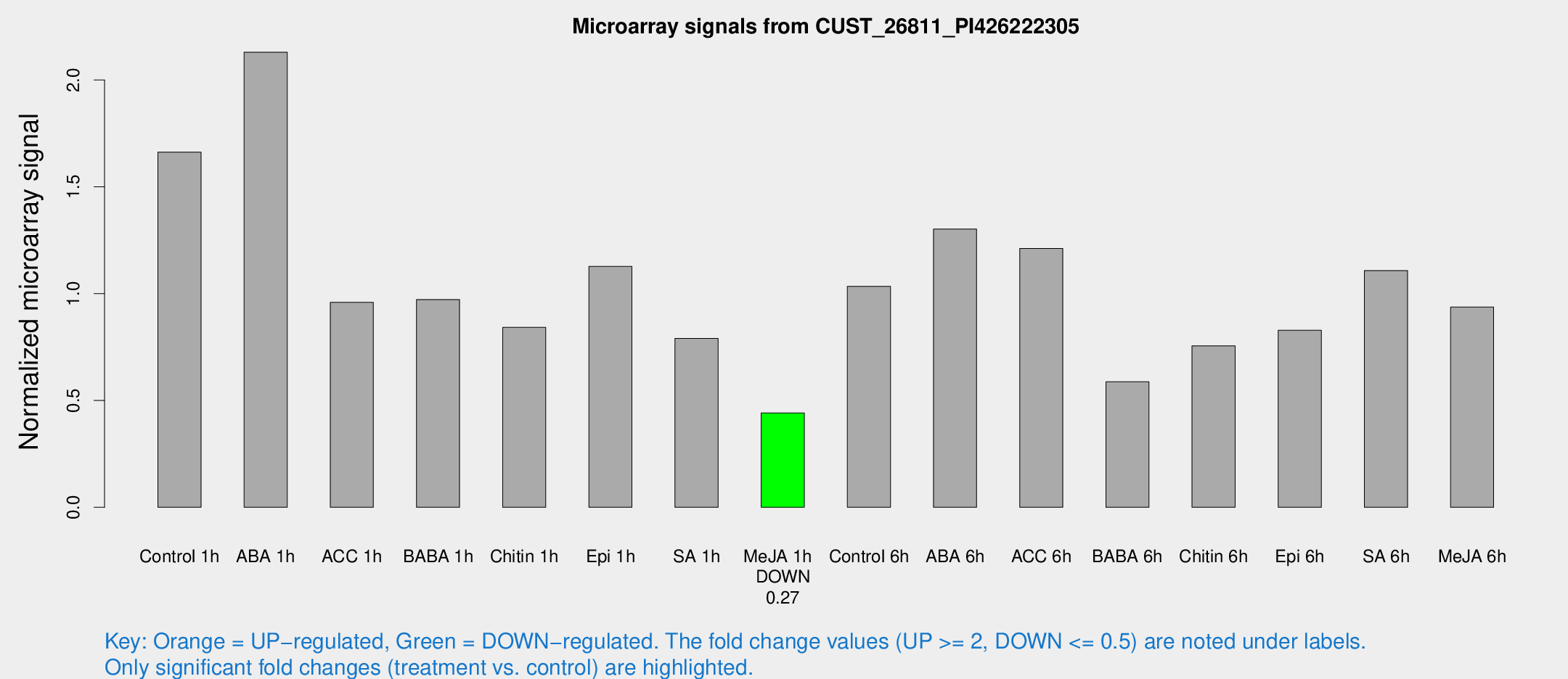
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1805.62 | 163.416 | 1.66257 | 0.0960417 |
| ABA 1h | 2059.09 | 239.71 | 2.13019 | 0.180607 |
| ACC 1h | 1181.01 | 338.552 | 0.959409 | 0.278142 |
| BABA 1h | 1115.18 | 308.062 | 0.972522 | 0.234278 |
| Chitin 1h | 839.457 | 148.873 | 0.842045 | 0.100786 |
| Epi 1h | 1056.16 | 75.1272 | 1.12754 | 0.0786729 |
| SA 1h | 933.602 | 212.151 | 0.790656 | 0.176886 |
| Me-JA 1h | 392.517 | 43.3287 | 0.441293 | 0.0257985 |
| Control 6h | 1164.54 | 250.093 | 1.03444 | 0.186858 |
| ABA 6h | 1518.05 | 192.971 | 1.30255 | 0.123411 |
| ACC 6h | 1497.1 | 86.5953 | 1.21132 | 0.155543 |
| BABA 6h | 719.264 | 90.7125 | 0.587626 | 0.0810449 |
| Chitin 6h | 890.465 | 152.754 | 0.756015 | 0.105117 |
| Epi 6h | 1022.91 | 134.036 | 0.828753 | 0.0490073 |
| SA 6h | 1256.02 | 305.547 | 1.1077 | 0.173733 |
| Me-JA 6h | 1011.09 | 77.8959 | 0.937215 | 0.102388 |
Source Transcript PGSC0003DMT400035281 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G21760.1 | +1 | 2e-52 | 179 | 86/169 (51%) | UDP-Glycosyltransferase superfamily protein | chr3:7667099-7668556 FORWARD LENGTH=485 |
