Probe CUST_26468_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26468_PI426222305 | JHI_St_60k_v1 | DMT400037143 | GAAACTGCAGCTAGAAGAAGATGAATTTGACACACACACACACAAAATAAGCTAAATGCA |
All Microarray Probes Designed to Gene DMG400014322
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26392_PI426222305 | JHI_St_60k_v1 | DMT400037142 | GAAACTGCAGCTAGAAGAAGATGAATTTGACACACACACACACAAAATAAGCTAAATGCA |
| CUST_26468_PI426222305 | JHI_St_60k_v1 | DMT400037143 | GAAACTGCAGCTAGAAGAAGATGAATTTGACACACACACACACAAAATAAGCTAAATGCA |
Microarray Signals from CUST_26468_PI426222305
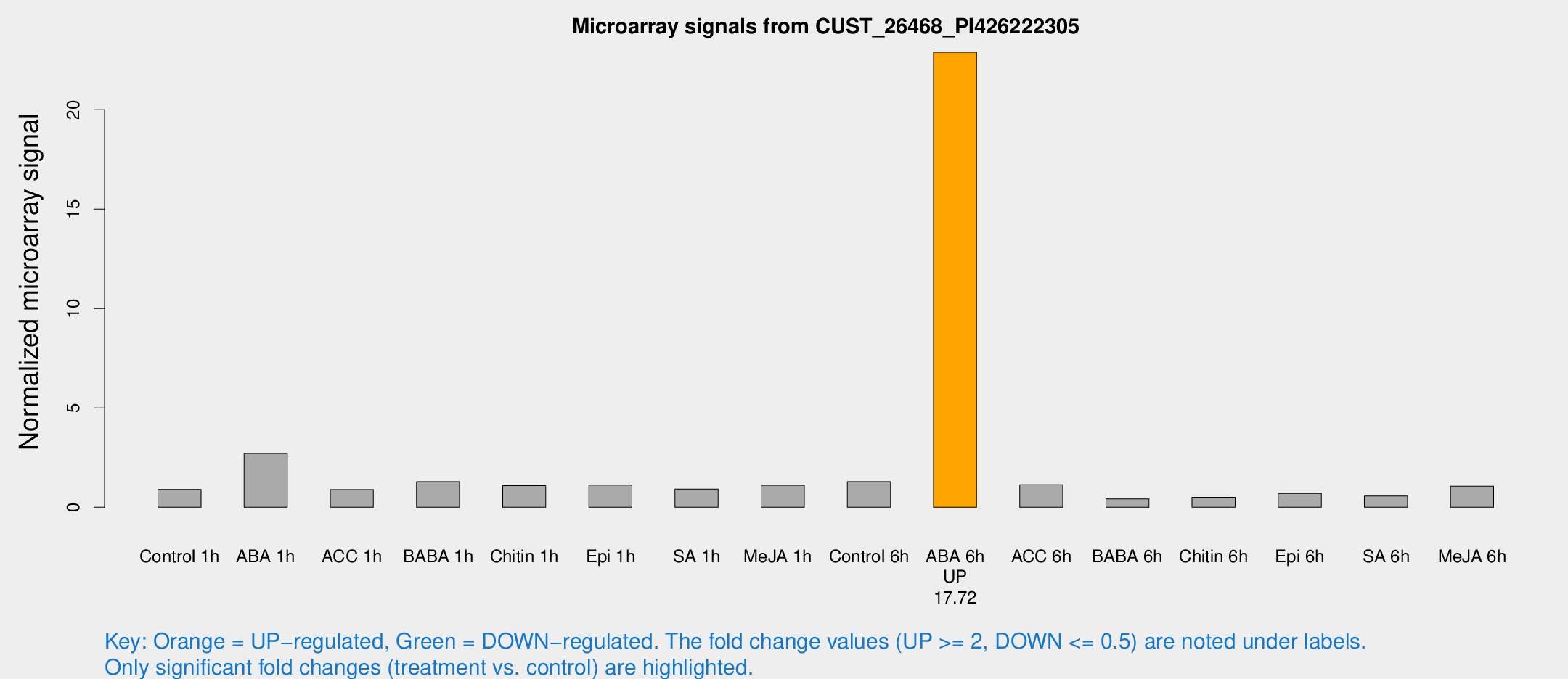
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 45.5532 | 12.1908 | 0.899427 | 0.311317 |
| ABA 1h | 118.354 | 28.1844 | 2.71556 | 0.815462 |
| ACC 1h | 51.0869 | 21.2792 | 0.889545 | 0.363816 |
| BABA 1h | 59.2127 | 8.29383 | 1.28489 | 0.113019 |
| Chitin 1h | 51.6529 | 16.2633 | 1.08458 | 0.425921 |
| Epi 1h | 47.1403 | 10.0148 | 1.11678 | 0.238403 |
| SA 1h | 47.3672 | 11.1934 | 0.917951 | 0.221753 |
| Me-JA 1h | 42.5237 | 4.3612 | 1.1104 | 0.146713 |
| Control 6h | 70.2594 | 27.4865 | 1.29138 | 0.564433 |
| ABA 6h | 1215.79 | 320.191 | 22.8889 | 4.63726 |
| ACC 6h | 62.542 | 10.7437 | 1.13058 | 0.355156 |
| BABA 6h | 25.4088 | 7.64612 | 0.425237 | 0.208656 |
| Chitin 6h | 24.9848 | 4.24958 | 0.500311 | 0.0856299 |
| Epi 6h | 37.3097 | 4.8004 | 0.699241 | 0.128138 |
| SA 6h | 28.7598 | 7.40573 | 0.571244 | 0.111519 |
| Me-JA 6h | 50.0173 | 6.25211 | 1.05686 | 0.11405 |
Source Transcript PGSC0003DMT400037143 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G62040.1 | +1 | 5e-82 | 249 | 120/169 (71%) | PEBP (phosphatidylethanolamine-binding protein) family protein | chr5:24922810-24923709 FORWARD LENGTH=177 |
