Probe CUST_26392_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26392_PI426222305 | JHI_St_60k_v1 | DMT400037142 | GAAACTGCAGCTAGAAGAAGATGAATTTGACACACACACACACAAAATAAGCTAAATGCA |
All Microarray Probes Designed to Gene DMG400014322
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26392_PI426222305 | JHI_St_60k_v1 | DMT400037142 | GAAACTGCAGCTAGAAGAAGATGAATTTGACACACACACACACAAAATAAGCTAAATGCA |
| CUST_26468_PI426222305 | JHI_St_60k_v1 | DMT400037143 | GAAACTGCAGCTAGAAGAAGATGAATTTGACACACACACACACAAAATAAGCTAAATGCA |
Microarray Signals from CUST_26392_PI426222305
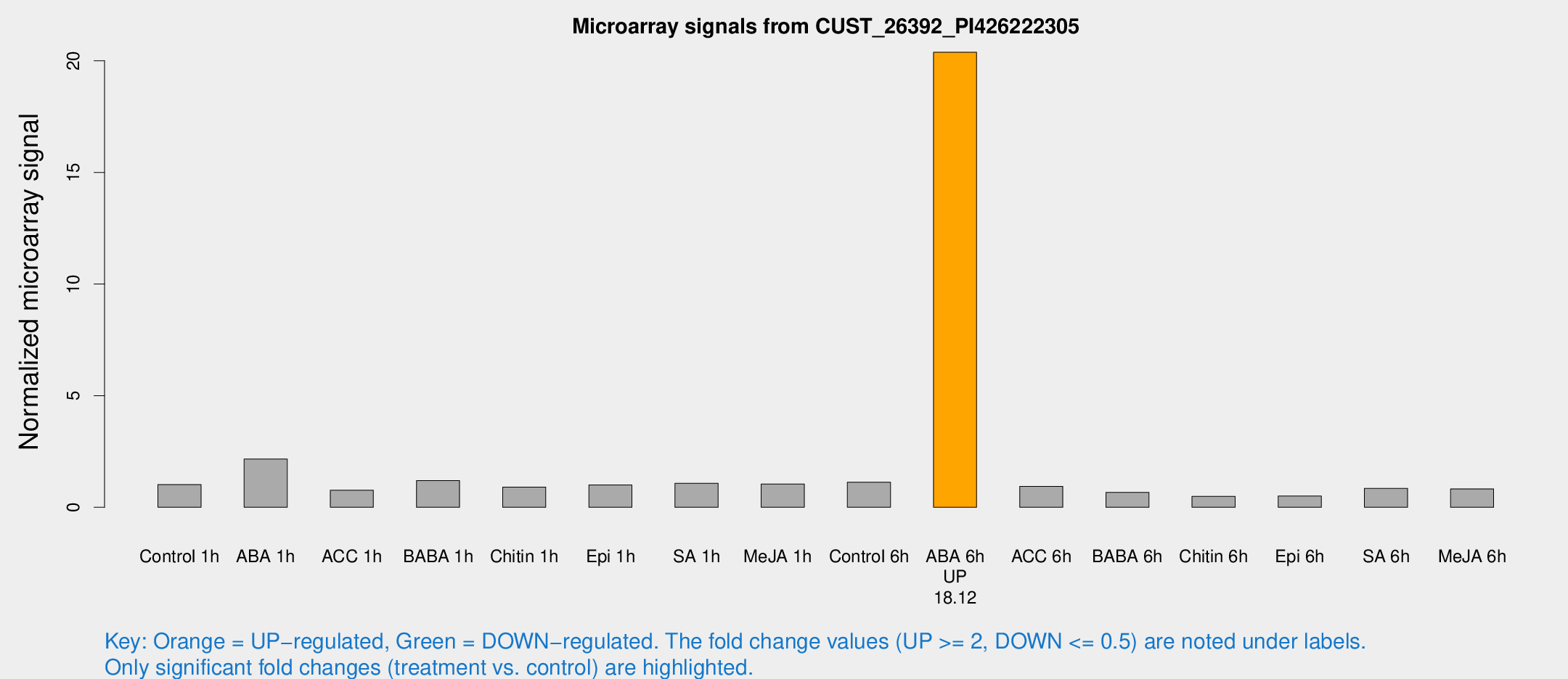
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 57.2294 | 13.5702 | 1.01662 | 0.298762 |
| ABA 1h | 105.311 | 20.2176 | 2.15932 | 0.536183 |
| ACC 1h | 46.6651 | 15.3018 | 0.76434 | 0.223526 |
| BABA 1h | 61.5664 | 5.21383 | 1.19748 | 0.101478 |
| Chitin 1h | 51.2272 | 17.0646 | 0.908033 | 0.461546 |
| Epi 1h | 49.1051 | 12.6357 | 1.00238 | 0.258487 |
| SA 1h | 59.424 | 5.15981 | 1.07719 | 0.0915366 |
| Me-JA 1h | 47.8817 | 10.0593 | 1.04498 | 0.344825 |
| Control 6h | 64.5552 | 18.1466 | 1.12463 | 0.303902 |
| ABA 6h | 1236.46 | 336.024 | 20.3837 | 4.24117 |
| ACC 6h | 57.8268 | 5.74779 | 0.939797 | 0.208571 |
| BABA 6h | 40.2681 | 4.72211 | 0.666447 | 0.0794388 |
| Chitin 6h | 28.6378 | 4.41077 | 0.491682 | 0.0999086 |
| Epi 6h | 31.5376 | 6.25904 | 0.506261 | 0.138195 |
| SA 6h | 44.5257 | 4.70545 | 0.843684 | 0.0894741 |
| Me-JA 6h | 45.8087 | 10.0952 | 0.82511 | 0.167542 |
Source Transcript PGSC0003DMT400037142 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G62040.1 | +1 | 4e-62 | 147 | 65/98 (66%) | PEBP (phosphatidylethanolamine-binding protein) family protein | chr5:24922810-24923709 FORWARD LENGTH=177 |
