Probe CUST_26368_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26368_PI426222305 | JHI_St_60k_v1 | DMT400037109 | GATGATGTTTTTCCTGGACTTGTCATTAAGATTGTTCCTTTCAACAATAACAAGTCATGG |
All Microarray Probes Designed to Gene DMG400014304
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26368_PI426222305 | JHI_St_60k_v1 | DMT400037109 | GATGATGTTTTTCCTGGACTTGTCATTAAGATTGTTCCTTTCAACAATAACAAGTCATGG |
| CUST_26384_PI426222305 | JHI_St_60k_v1 | DMT400037110 | TAGCTTCATGGATATGTGTAATTAGCCTCATGGCACTTGTATTAATGAGTTCTGTTAGGG |
Microarray Signals from CUST_26368_PI426222305
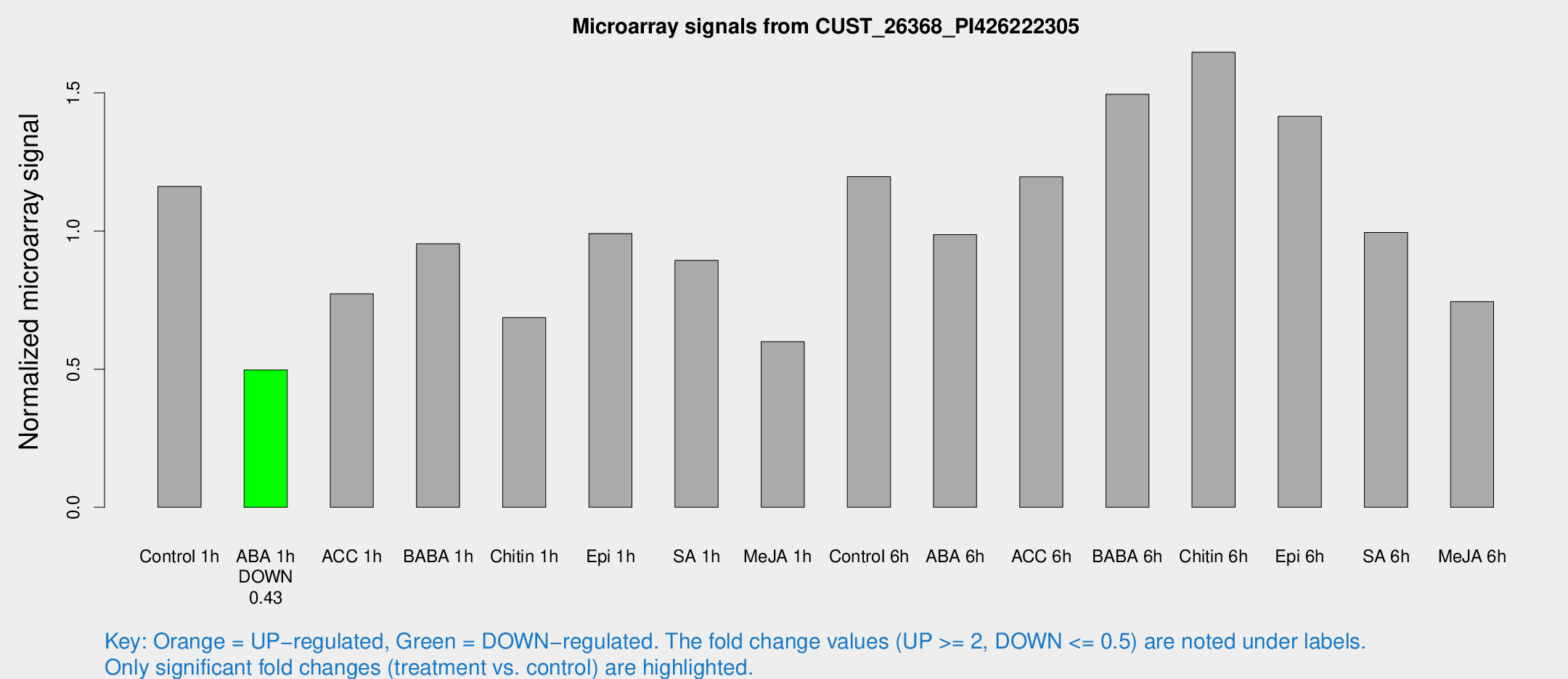
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 285.589 | 16.8623 | 1.16166 | 0.0684994 |
| ABA 1h | 113.243 | 23.5827 | 0.496856 | 0.0804194 |
| ACC 1h | 251.533 | 96.5943 | 0.772677 | 0.461949 |
| BABA 1h | 229.691 | 28.852 | 0.953844 | 0.0572326 |
| Chitin 1h | 151.736 | 9.40226 | 0.686948 | 0.0425289 |
| Epi 1h | 210.893 | 12.6413 | 0.991164 | 0.0593681 |
| SA 1h | 227.789 | 23.5547 | 0.893962 | 0.0684377 |
| Me-JA 1h | 123.018 | 19.5388 | 0.599311 | 0.0657089 |
| Control 6h | 315.998 | 74.3618 | 1.19694 | 0.230689 |
| ABA 6h | 262.802 | 37.5492 | 0.986643 | 0.178713 |
| ACC 6h | 348.261 | 64.6218 | 1.1963 | 0.10685 |
| BABA 6h | 442.631 | 107.5 | 1.49527 | 0.388362 |
| Chitin 6h | 435.168 | 45.9185 | 1.64737 | 0.164513 |
| Epi 6h | 396.042 | 39.7835 | 1.41558 | 0.259011 |
| SA 6h | 251.439 | 52.6232 | 0.994818 | 0.164658 |
| Me-JA 6h | 198.405 | 52.6541 | 0.744842 | 0.172254 |
Source Transcript PGSC0003DMT400037109 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G62150.1 | +1 | 2e-37 | 124 | 59/86 (69%) | peptidoglycan-binding LysM domain-containing protein | chr5:24958325-24958633 FORWARD LENGTH=102 |
