Probe CUST_24989_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24989_PI426222305 | JHI_St_60k_v1 | DMT400084733 | TTCTCACCGTCTTAATCAGCTTCGTACTAGTTACTGGAATCATACAGCTCTGTCAAAGAG |
All Microarray Probes Designed to Gene DMG400034310
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24989_PI426222305 | JHI_St_60k_v1 | DMT400084733 | TTCTCACCGTCTTAATCAGCTTCGTACTAGTTACTGGAATCATACAGCTCTGTCAAAGAG |
| CUST_24984_PI426222305 | JHI_St_60k_v1 | DMT400084734 | ATTCCAAAAGCCCAAAGGTTATTGTAGCAGGTGTAGCACGCCGCTTTTGTCAACAGTGTA |
Microarray Signals from CUST_24989_PI426222305
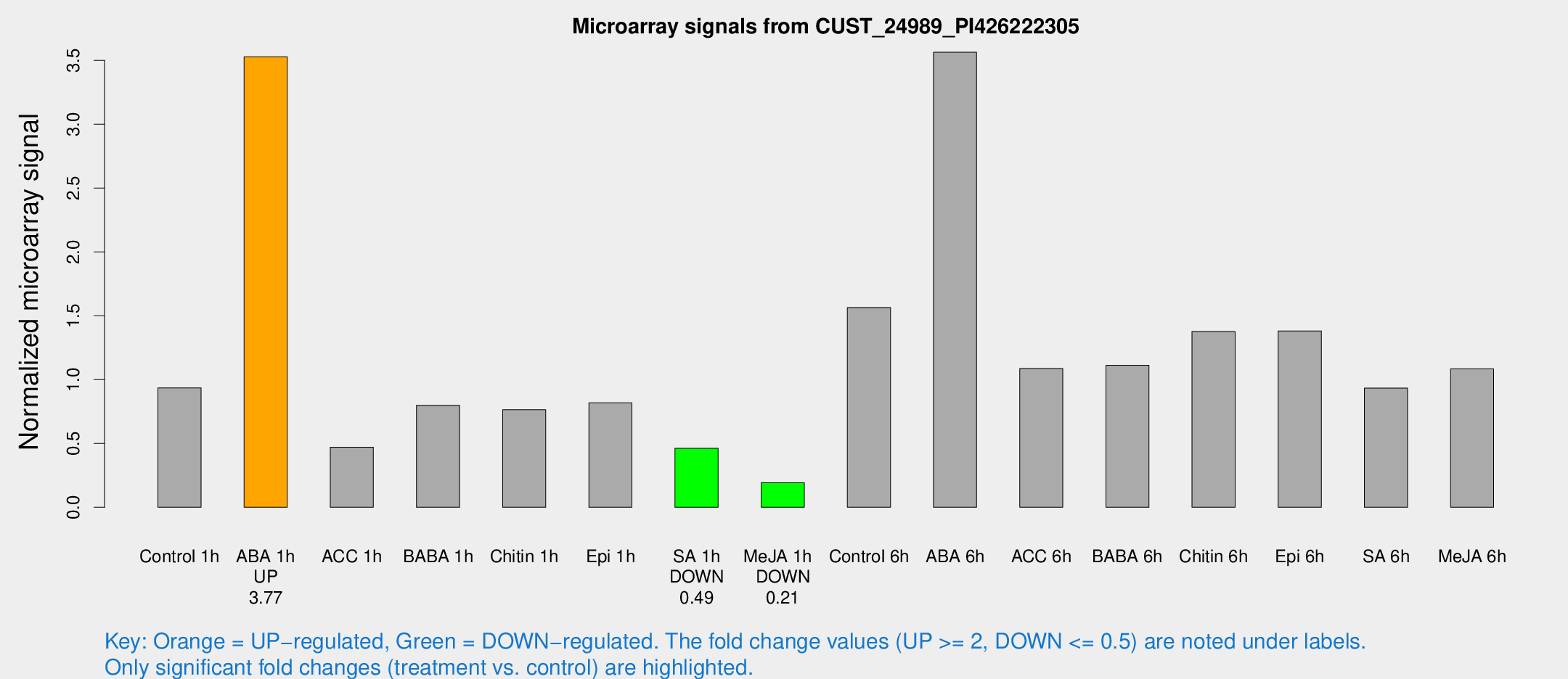
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 52.3875 | 7.55369 | 0.935758 | 0.131147 |
| ABA 1h | 174.935 | 25.5476 | 3.52827 | 0.677254 |
| ACC 1h | 28.6801 | 7.04761 | 0.471121 | 0.104056 |
| BABA 1h | 42.8788 | 4.99045 | 0.798793 | 0.0822053 |
| Chitin 1h | 42.2009 | 12.8344 | 0.764475 | 0.249725 |
| Epi 1h | 39.3527 | 4.31151 | 0.818411 | 0.0896449 |
| SA 1h | 26.4168 | 3.54176 | 0.461595 | 0.0637296 |
| Me-JA 1h | 9.27835 | 3.35091 | 0.192287 | 0.0782461 |
| Control 6h | 89.5901 | 16.5081 | 1.56439 | 0.184328 |
| ABA 6h | 211.691 | 26.7196 | 3.56416 | 0.329816 |
| ACC 6h | 71.9794 | 15.8304 | 1.08652 | 0.129527 |
| BABA 6h | 72.1881 | 16.378 | 1.11317 | 0.240214 |
| Chitin 6h | 80.8765 | 6.31288 | 1.37609 | 0.148972 |
| Epi 6h | 88.5466 | 15.5827 | 1.38168 | 0.360021 |
| SA 6h | 71.8218 | 32.1927 | 0.932987 | 1.17464 |
| Me-JA 6h | 67.124 | 23.6532 | 1.08381 | 0.309705 |
Source Transcript PGSC0003DMT400084733 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G43270.2 | +3 | 5e-54 | 197 | 137/321 (43%) | squamosa promoter binding protein-like 2 | chr5:17360527-17362143 REVERSE LENGTH=419 |
