Probe CUST_24597_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24597_PI426222305 | JHI_St_60k_v1 | DMT400054479 | GAGATAATTGCTTGTCTGCTGACCTCCTTTTTGTGAACATCGTGCCTTTATTACTTTTTC |
All Microarray Probes Designed to Gene DMG400021143
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24597_PI426222305 | JHI_St_60k_v1 | DMT400054479 | GAGATAATTGCTTGTCTGCTGACCTCCTTTTTGTGAACATCGTGCCTTTATTACTTTTTC |
Microarray Signals from CUST_24597_PI426222305
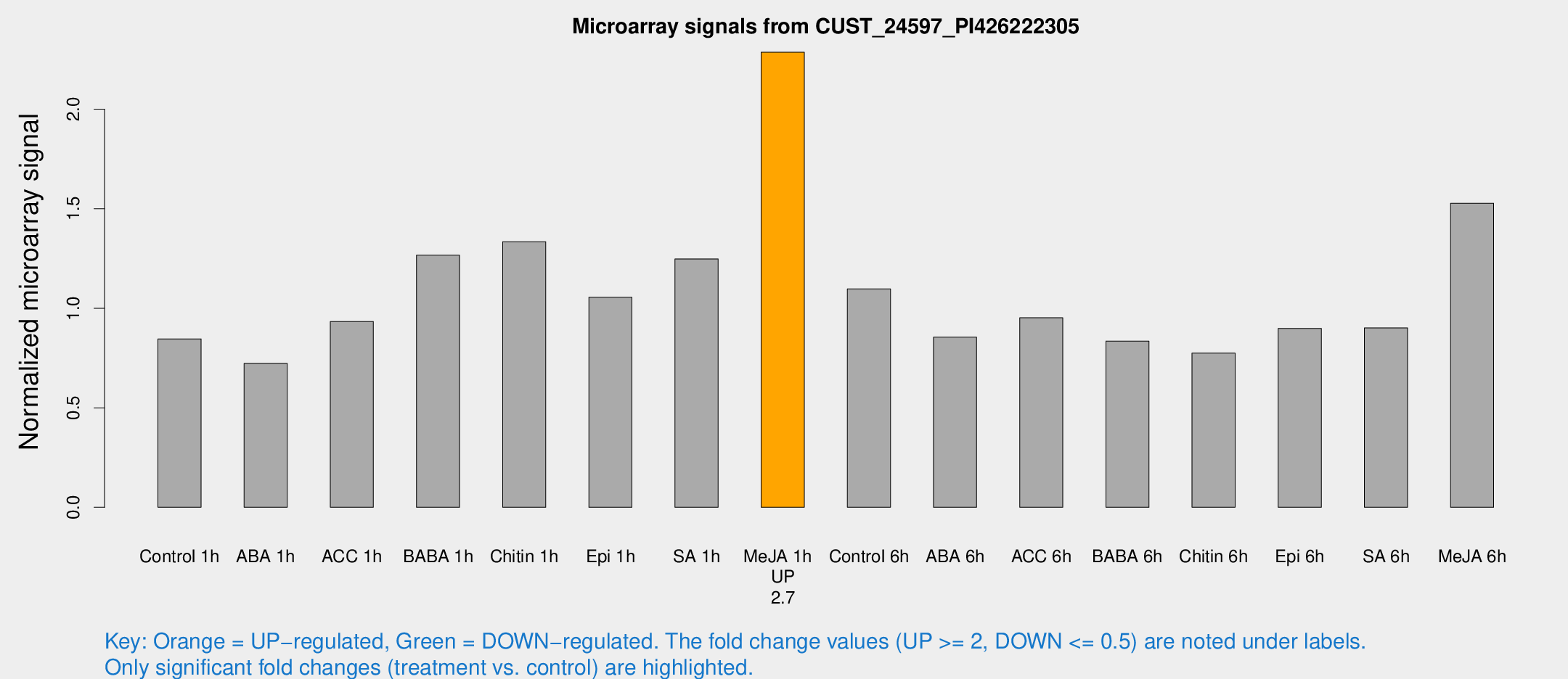
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 537.368 | 93.5389 | 0.846252 | 0.159328 |
| ABA 1h | 397.893 | 47.3938 | 0.722442 | 0.0739171 |
| ACC 1h | 601.023 | 85.7022 | 0.93347 | 0.0824587 |
| BABA 1h | 760.421 | 92.7514 | 1.26635 | 0.0786835 |
| Chitin 1h | 736.966 | 42.6828 | 1.33442 | 0.0898532 |
| Epi 1h | 563.534 | 40.1063 | 1.05536 | 0.0735976 |
| SA 1h | 804.415 | 118.356 | 1.24805 | 0.159638 |
| Me-JA 1h | 1160.73 | 127.073 | 2.28635 | 0.13215 |
| Control 6h | 708.862 | 143.695 | 1.09696 | 0.149883 |
| ABA 6h | 564.722 | 60.7856 | 0.854694 | 0.0568411 |
| ACC 6h | 670.503 | 38.9019 | 0.952174 | 0.12212 |
| BABA 6h | 573.821 | 33.322 | 0.834863 | 0.0484607 |
| Chitin 6h | 509.863 | 43.5657 | 0.774965 | 0.0674704 |
| Epi 6h | 644.124 | 126.609 | 0.898317 | 0.170454 |
| SA 6h | 551.385 | 48.1866 | 0.901194 | 0.0528018 |
| Me-JA 6h | 934.718 | 54.1159 | 1.52757 | 0.10704 |
Source Transcript PGSC0003DMT400054479 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G38970.1 | +1 | 8e-172 | 518 | 304/653 (47%) | Zinc finger (C3HC4-type RING finger) family protein | chr2:16274135-16276651 FORWARD LENGTH=692 |
