Probe CUST_24523_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24523_PI426222305 | JHI_St_60k_v1 | DMT400054403 | CATCTTCTGTTCCCCAAAAAAGGAGTTAAAATGGAAGTCGTCGCAATCGAATTTTTATTT |
All Microarray Probes Designed to Gene DMG400021116
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24523_PI426222305 | JHI_St_60k_v1 | DMT400054403 | CATCTTCTGTTCCCCAAAAAAGGAGTTAAAATGGAAGTCGTCGCAATCGAATTTTTATTT |
Microarray Signals from CUST_24523_PI426222305
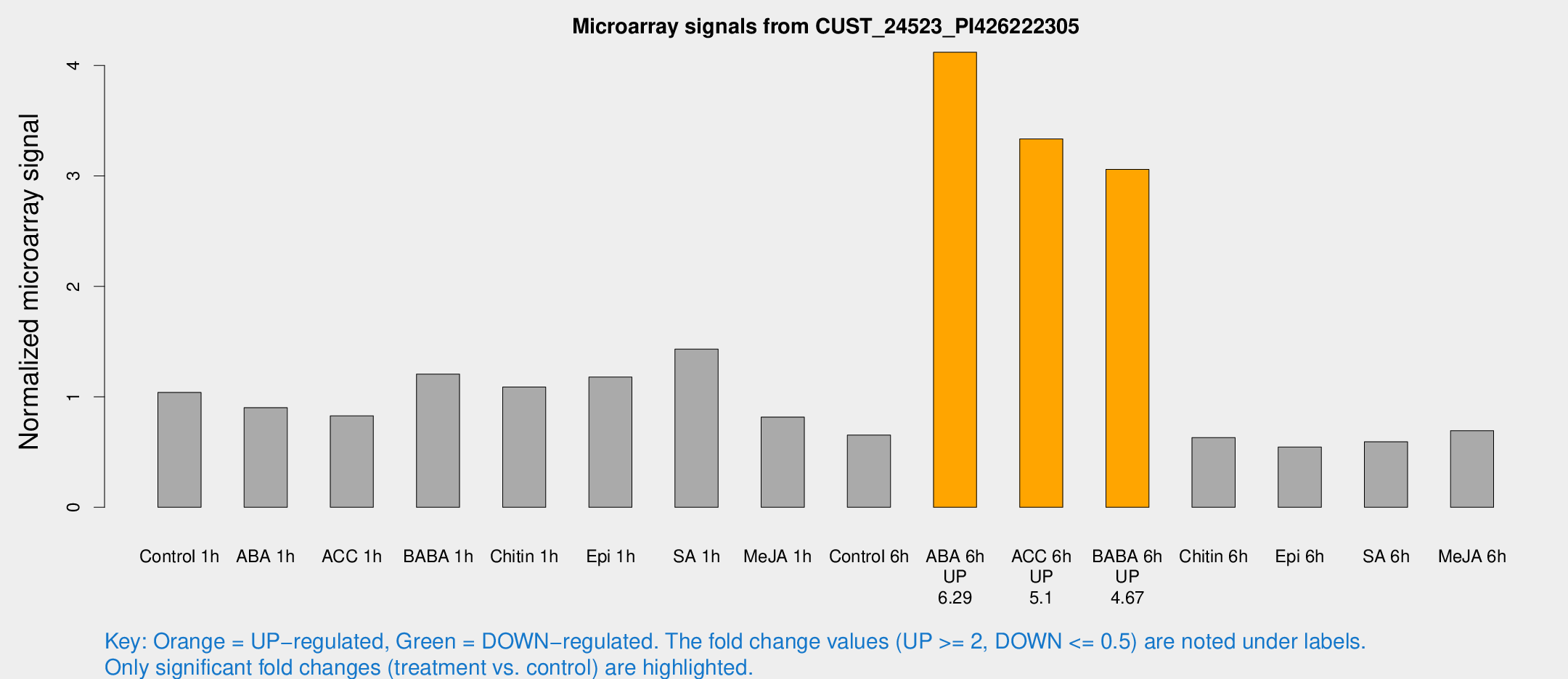
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 111.654 | 7.28671 | 1.04027 | 0.135452 |
| ABA 1h | 91.967 | 22.3374 | 0.902342 | 0.213934 |
| ACC 1h | 100.074 | 28.1826 | 0.828102 | 0.220103 |
| BABA 1h | 140.232 | 46.2328 | 1.20639 | 0.335515 |
| Chitin 1h | 107.698 | 18.5994 | 1.08871 | 0.142152 |
| Epi 1h | 109.224 | 7.11033 | 1.17911 | 0.0767708 |
| SA 1h | 168.469 | 39.2415 | 1.43301 | 0.323807 |
| Me-JA 1h | 73.5145 | 13.0302 | 0.817104 | 0.071908 |
| Control 6h | 76.1118 | 21.6768 | 0.65469 | 0.152517 |
| ABA 6h | 534.655 | 172.501 | 4.12064 | 1.4936 |
| ACC 6h | 413.784 | 44.8469 | 3.33634 | 0.782454 |
| BABA 6h | 387.75 | 96.605 | 3.05991 | 0.667968 |
| Chitin 6h | 72.6098 | 7.37841 | 0.631583 | 0.0498476 |
| Epi 6h | 75.4254 | 29.2465 | 0.545346 | 0.181304 |
| SA 6h | 70.645 | 21.4647 | 0.593098 | 0.155657 |
| Me-JA 6h | 74.4212 | 6.05383 | 0.694142 | 0.0513931 |
Source Transcript PGSC0003DMT400054403 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G49690.1 | +1 | 0.0 | 540 | 265/474 (56%) | UDP-Glycosyltransferase superfamily protein | chr5:20189968-20191350 REVERSE LENGTH=460 |
