Probe CUST_23856_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23856_PI426222305 | JHI_St_60k_v1 | DMT400076587 | TCGAATCCTTGGTTTTACTTCACCTGGTGTAGAGGTATCAGTGGATTGTGCAATAGTGAT |
All Microarray Probes Designed to Gene DMG400029777
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23856_PI426222305 | JHI_St_60k_v1 | DMT400076587 | TCGAATCCTTGGTTTTACTTCACCTGGTGTAGAGGTATCAGTGGATTGTGCAATAGTGAT |
| CUST_24036_PI426222305 | JHI_St_60k_v1 | DMT400076586 | TGTGAATTTAACAGTACATGATGAATTTGAGCAGGTATCAGTGGATTGTGCAATAGTGAT |
Microarray Signals from CUST_23856_PI426222305
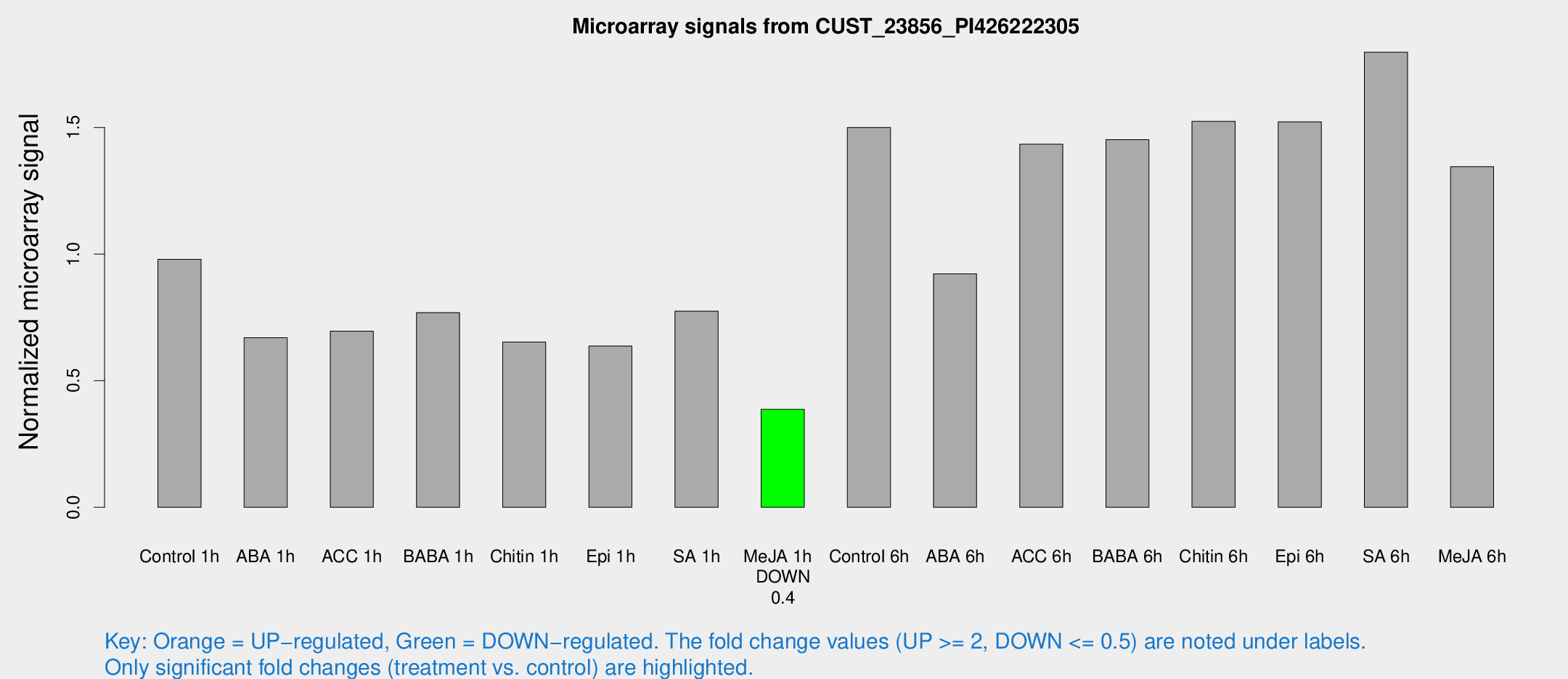
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 832.503 | 118.285 | 0.97885 | 0.0789208 |
| ABA 1h | 495.277 | 28.8244 | 0.670094 | 0.0388831 |
| ACC 1h | 653.535 | 179.601 | 0.695678 | 0.177065 |
| BABA 1h | 648.758 | 144.173 | 0.768614 | 0.111807 |
| Chitin 1h | 494.912 | 54.6701 | 0.652586 | 0.037882 |
| Epi 1h | 469.98 | 73.7877 | 0.636742 | 0.100547 |
| SA 1h | 675.942 | 90.7789 | 0.774352 | 0.0789675 |
| Me-JA 1h | 267.269 | 34.9718 | 0.386651 | 0.0227685 |
| Control 6h | 1301.64 | 248.131 | 1.49945 | 0.194607 |
| ABA 6h | 817.698 | 47.3951 | 0.921937 | 0.0533483 |
| ACC 6h | 1372.93 | 79.5259 | 1.43383 | 0.138401 |
| BABA 6h | 1367.83 | 143.622 | 1.45184 | 0.157125 |
| Chitin 6h | 1351.65 | 78.1295 | 1.5242 | 0.0880821 |
| Epi 6h | 1436.98 | 108.848 | 1.52161 | 0.120779 |
| SA 6h | 1504.46 | 196.228 | 1.79691 | 0.171602 |
| Me-JA 6h | 1148.38 | 194.393 | 1.34501 | 0.116988 |
Source Transcript PGSC0003DMT400076587 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G01080.1 | +3 | 5e-64 | 208 | 119/165 (72%) | Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family | chr2:78038-79176 FORWARD LENGTH=231 |
