Probe CUST_22762_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22762_PI426222305 | JHI_St_60k_v1 | DMT400078035 | GTGCGCACCAGAAAGTAGGTTAAAATTGATCCTTCTACCCTGTAAATTTGTTAATTGATA |
All Microarray Probes Designed to Gene DMG400030352
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22543_PI426222305 | JHI_St_60k_v1 | DMT400078034 | GTGCGCACCAGAAAGTAGGTTAAAATTGATCCTTCTACCCTGTAAATTTGTTAATTGATA |
| CUST_22762_PI426222305 | JHI_St_60k_v1 | DMT400078035 | GTGCGCACCAGAAAGTAGGTTAAAATTGATCCTTCTACCCTGTAAATTTGTTAATTGATA |
Microarray Signals from CUST_22762_PI426222305
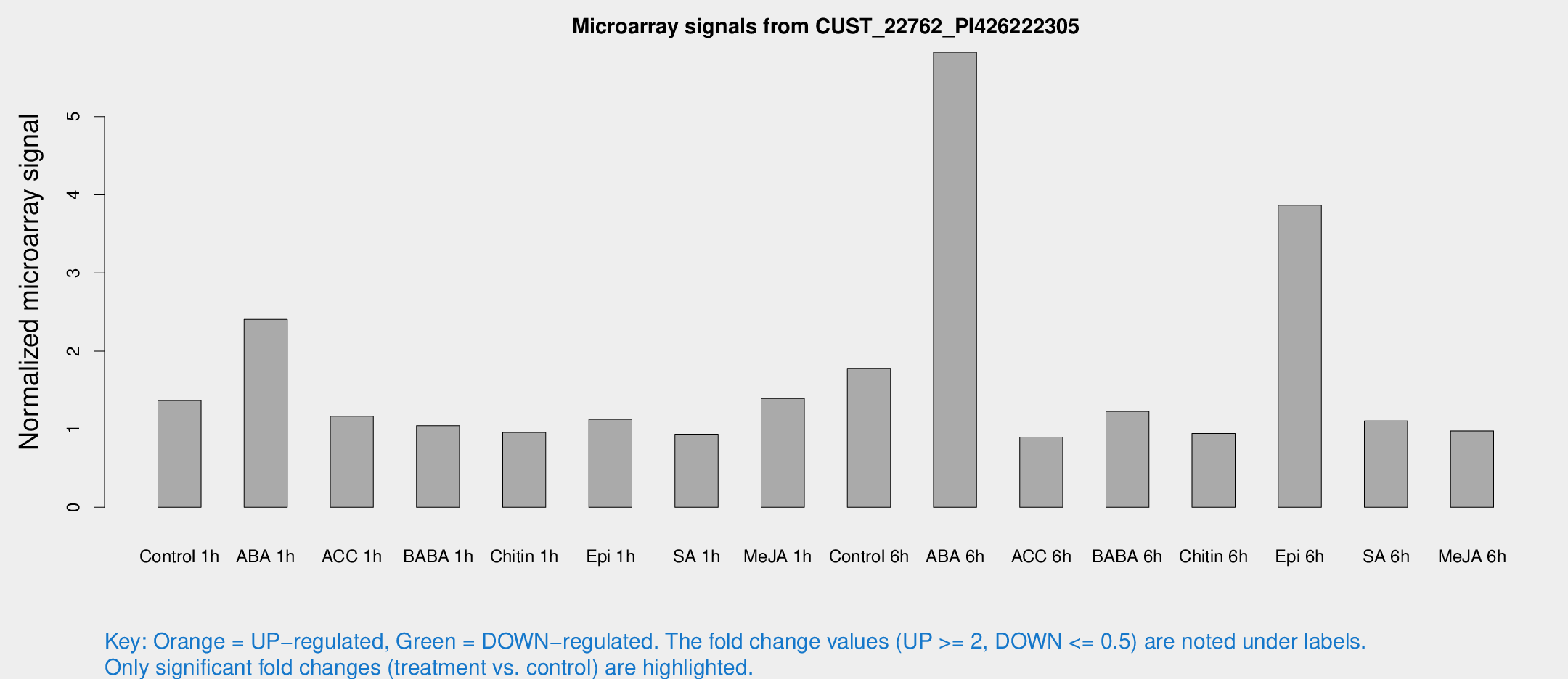
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11.9405 | 5.54767 | 1.36718 | 0.812978 |
| ABA 1h | 17.951 | 6.16093 | 2.40575 | 1.44409 |
| ACC 1h | 8.81522 | 4.83092 | 1.16494 | 0.60842 |
| BABA 1h | 7.30137 | 3.7708 | 1.04458 | 0.55247 |
| Chitin 1h | 6.18768 | 3.59527 | 0.95935 | 0.556256 |
| Epi 1h | 7.0854 | 3.52284 | 1.12688 | 0.579876 |
| SA 1h | 6.97781 | 3.59577 | 0.936878 | 0.494634 |
| Me-JA 1h | 8.68802 | 3.70542 | 1.39298 | 0.675591 |
| Control 6h | 16.1227 | 7.80889 | 1.77778 | 1.04118 |
| ABA 6h | 46.6735 | 10.3201 | 5.82458 | 1.25077 |
| ACC 6h | 7.5728 | 4.50876 | 0.898745 | 0.521089 |
| BABA 6h | 11.5131 | 4.76181 | 1.22878 | 0.575803 |
| Chitin 6h | 7.21288 | 4.17841 | 0.946147 | 0.547905 |
| Epi 6h | 86.4363 | 73.3452 | 3.86811 | 10.5422 |
| SA 6h | 7.91234 | 3.94853 | 1.10595 | 0.559671 |
| Me-JA 6h | 7.0713 | 3.69638 | 0.978946 | 0.52147 |
Source Transcript PGSC0003DMT400078035 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G58800.1 | +3 | 2e-95 | 290 | 154/193 (80%) | Quinone reductase family protein | chr5:23746032-23746895 REVERSE LENGTH=207 |
