Probe CUST_22702_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22702_PI426222305 | JHI_St_60k_v1 | DMT400078286 | GGAAGAATCATGTATGCACTTTCAGTTTGGGAGCCTATTCATAAATAAAATCTTGCATTC |
All Microarray Probes Designed to Gene DMG400030468
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22702_PI426222305 | JHI_St_60k_v1 | DMT400078286 | GGAAGAATCATGTATGCACTTTCAGTTTGGGAGCCTATTCATAAATAAAATCTTGCATTC |
Microarray Signals from CUST_22702_PI426222305
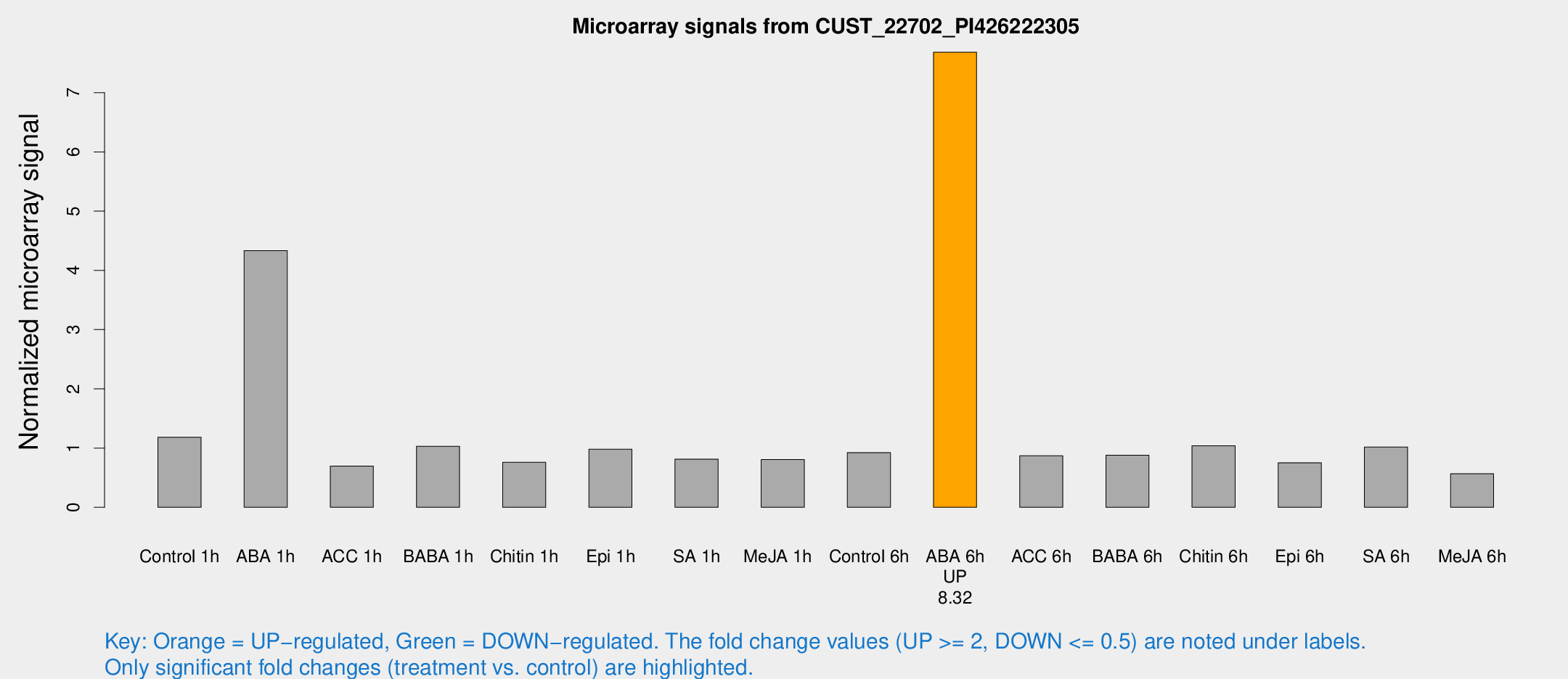
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 38.4876 | 3.90505 | 1.18336 | 0.133224 |
| ABA 1h | 135.225 | 35.8813 | 4.33358 | 1.06542 |
| ACC 1h | 25.4879 | 6.91254 | 0.695808 | 0.183722 |
| BABA 1h | 34.2063 | 7.45533 | 1.03097 | 0.156897 |
| Chitin 1h | 22.1945 | 3.53074 | 0.75919 | 0.120158 |
| Epi 1h | 28.8426 | 5.73128 | 0.980244 | 0.216331 |
| SA 1h | 27.2425 | 3.56078 | 0.813632 | 0.106527 |
| Me-JA 1h | 22.5748 | 5.18937 | 0.80551 | 0.140987 |
| Control 6h | 31.2326 | 5.86639 | 0.923838 | 0.129118 |
| ABA 6h | 269.266 | 35.3731 | 7.68286 | 0.621429 |
| ACC 6h | 34.1208 | 8.19851 | 0.869902 | 0.118359 |
| BABA 6h | 34.4851 | 10.0725 | 0.878769 | 0.251398 |
| Chitin 6h | 36.3994 | 4.43335 | 1.03815 | 0.147387 |
| Epi 6h | 30.8247 | 8.85743 | 0.751364 | 0.376378 |
| SA 6h | 36.2721 | 10.8457 | 1.01639 | 0.438097 |
| Me-JA 6h | 24.3731 | 10.4138 | 0.56644 | 0.378362 |
Source Transcript PGSC0003DMT400078286 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G37650.1 | +3 | 7e-152 | 462 | 240/417 (58%) | GRAS family transcription factor | chr2:15792623-15794779 FORWARD LENGTH=718 |
