Probe CUST_22576_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22576_PI426222305 | JHI_St_60k_v1 | DMT400078244 | CTCAAGTGGATCGTAGATCCTTTGTAGCAATGCAAATTAAATCCTTCCTTGATTAATGAT |
All Microarray Probes Designed to Gene DMG400030450
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22576_PI426222305 | JHI_St_60k_v1 | DMT400078244 | CTCAAGTGGATCGTAGATCCTTTGTAGCAATGCAAATTAAATCCTTCCTTGATTAATGAT |
| CUST_22827_PI426222305 | JHI_St_60k_v1 | DMT400078243 | CTCAAGTGGATCGTAGATCCTTTGTAGCAATGCAAATTAAATCCTTCCTTGATTAATGAT |
Microarray Signals from CUST_22576_PI426222305
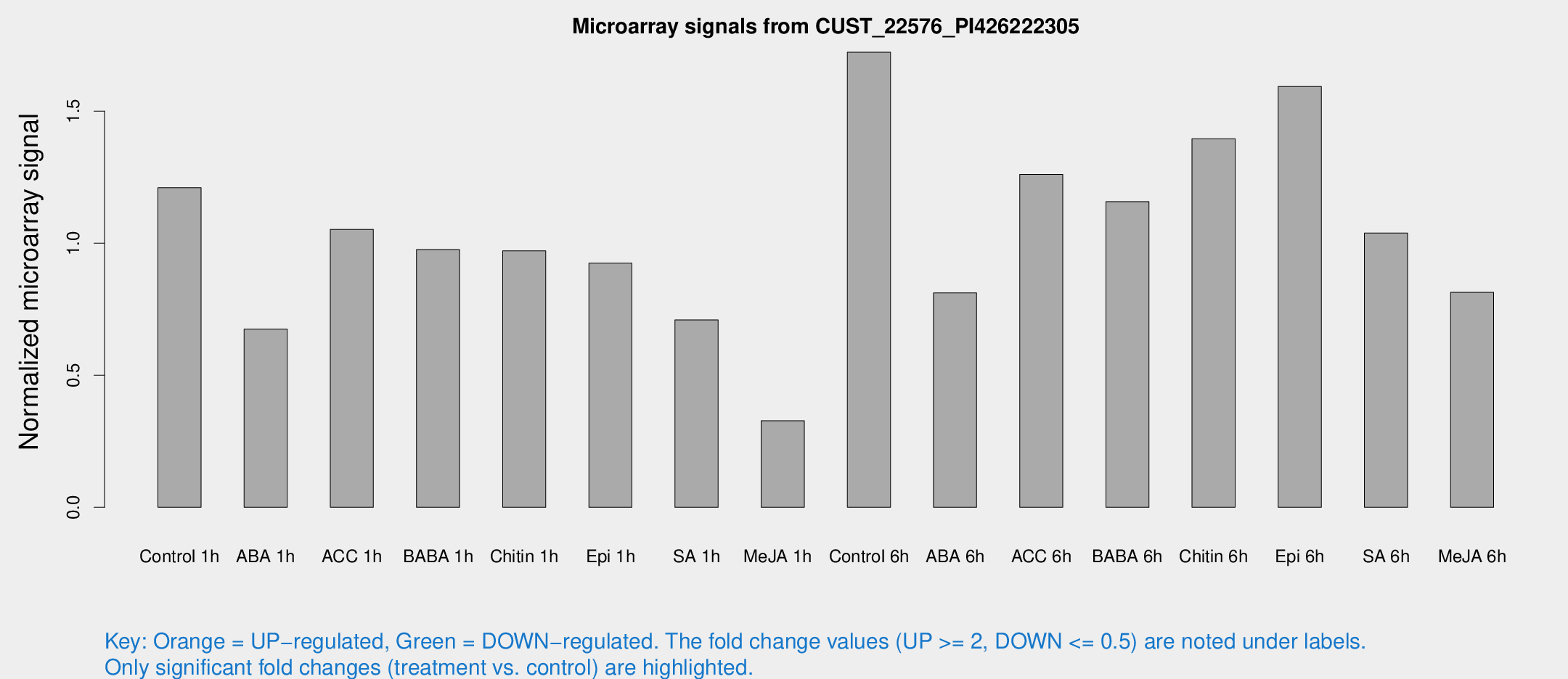
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 374.716 | 61.8845 | 1.21017 | 0.1336 |
| ABA 1h | 182.217 | 22.7681 | 0.674551 | 0.0665385 |
| ACC 1h | 350.334 | 93.4288 | 1.05206 | 0.223651 |
| BABA 1h | 289.542 | 44.6183 | 0.975707 | 0.0846187 |
| Chitin 1h | 277.385 | 60.1066 | 0.970894 | 0.167682 |
| Epi 1h | 244.391 | 31.7677 | 0.924149 | 0.0915788 |
| SA 1h | 228.602 | 43.5744 | 0.709457 | 0.111448 |
| Me-JA 1h | 82.625 | 13.0317 | 0.327389 | 0.0575751 |
| Control 6h | 566.621 | 155.696 | 1.7229 | 0.378684 |
| ABA 6h | 259.93 | 15.4856 | 0.81163 | 0.0482731 |
| ACC 6h | 444.815 | 67.8739 | 1.26043 | 0.0738458 |
| BABA 6h | 398.915 | 58.0615 | 1.15729 | 0.144519 |
| Chitin 6h | 448.887 | 32.6896 | 1.39519 | 0.118394 |
| Epi 6h | 574.413 | 127.033 | 1.59347 | 0.566555 |
| SA 6h | 319.733 | 61.7493 | 1.03813 | 0.145445 |
| Me-JA 6h | 262.415 | 63.3775 | 0.813979 | 0.173009 |
Source Transcript PGSC0003DMT400078244 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G58520.1 | +1 | 0.0 | 642 | 312/451 (69%) | Protein kinase superfamily protein | chr5:23655312-23657943 FORWARD LENGTH=604 |
