Probe CUST_22568_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22568_PI426222305 | JHI_St_60k_v1 | DMT400078202 | TCTGTTGGAGATTTGAAGTTGCTGTGCTCTCTGTATTCAAGTGCAATAGGAAATAAAATA |
All Microarray Probes Designed to Gene DMG400030427
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22568_PI426222305 | JHI_St_60k_v1 | DMT400078202 | TCTGTTGGAGATTTGAAGTTGCTGTGCTCTCTGTATTCAAGTGCAATAGGAAATAAAATA |
Microarray Signals from CUST_22568_PI426222305
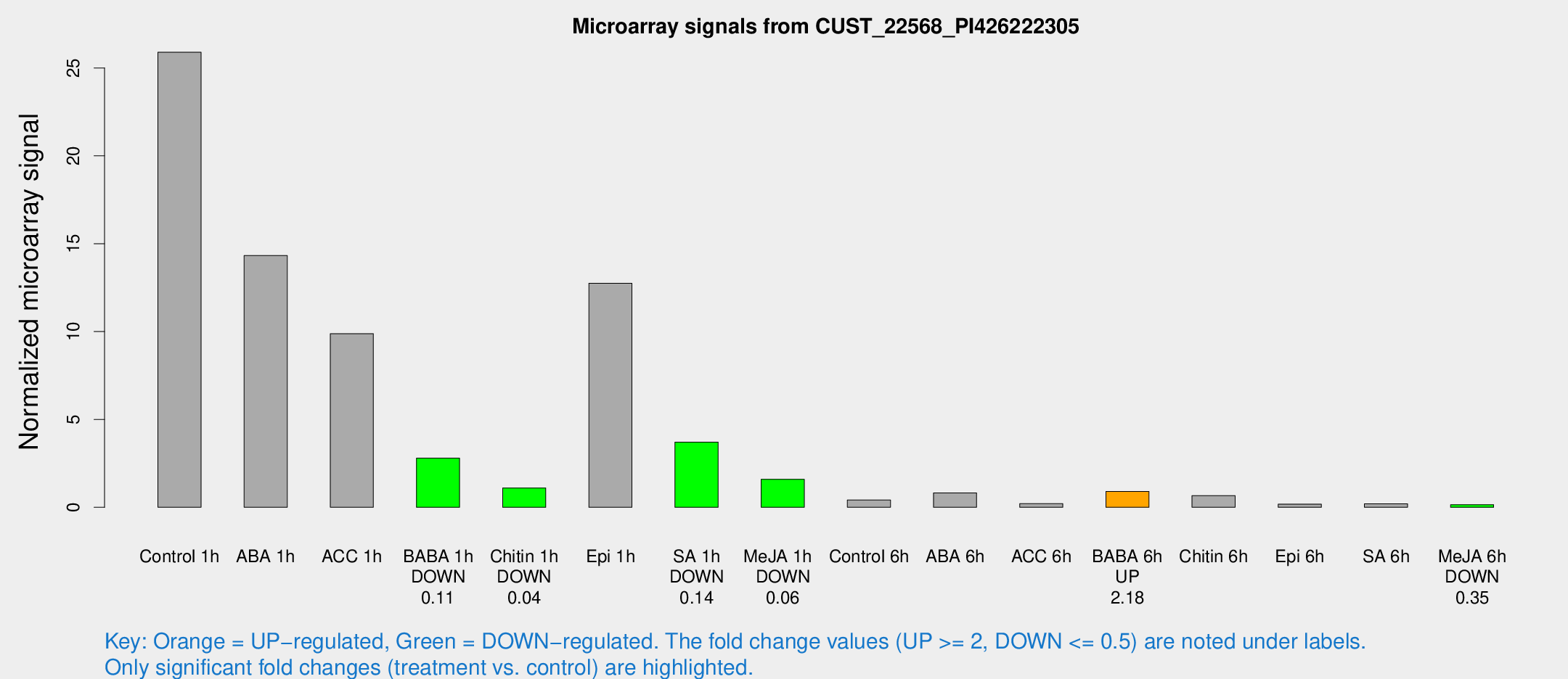
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1868.76 | 210.044 | 25.8937 | 2.89175 |
| ABA 1h | 2099.15 | 1693.28 | 14.3284 | 32.7 |
| ACC 1h | 1639.22 | 1145.13 | 9.8757 | 23.6147 |
| BABA 1h | 272.981 | 150.056 | 2.79702 | 1.83004 |
| Chitin 1h | 83.7073 | 35.0457 | 1.09947 | 0.442664 |
| Epi 1h | 963.011 | 409.094 | 12.747 | 7.42602 |
| SA 1h | 335.728 | 123.245 | 3.7128 | 2.02382 |
| Me-JA 1h | 93.8771 | 10.4131 | 1.59381 | 0.317811 |
| Control 6h | 30.8155 | 6.13162 | 0.41589 | 0.0753311 |
| ABA 6h | 63.2038 | 8.40031 | 0.819854 | 0.0699359 |
| ACC 6h | 20.1423 | 8.7103 | 0.20745 | 0.11973 |
| BABA 6h | 72.3592 | 5.97455 | 0.905065 | 0.0752535 |
| Chitin 6h | 52.6534 | 9.98571 | 0.666318 | 0.160014 |
| Epi 6h | 15.5972 | 4.58408 | 0.181326 | 0.0675853 |
| SA 6h | 16.3271 | 6.37378 | 0.197208 | 0.0722393 |
| Me-JA 6h | 10.3922 | 3.88102 | 0.146237 | 0.0547856 |
Source Transcript PGSC0003DMT400078202 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G07400.1 | +3 | 2e-68 | 211 | 118/157 (75%) | HSP20-like chaperones superfamily protein | chr1:2275148-2275621 FORWARD LENGTH=157 |
