Probe CUST_22543_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22543_PI426222305 | JHI_St_60k_v1 | DMT400078034 | GTGCGCACCAGAAAGTAGGTTAAAATTGATCCTTCTACCCTGTAAATTTGTTAATTGATA |
All Microarray Probes Designed to Gene DMG400030352
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22543_PI426222305 | JHI_St_60k_v1 | DMT400078034 | GTGCGCACCAGAAAGTAGGTTAAAATTGATCCTTCTACCCTGTAAATTTGTTAATTGATA |
| CUST_22762_PI426222305 | JHI_St_60k_v1 | DMT400078035 | GTGCGCACCAGAAAGTAGGTTAAAATTGATCCTTCTACCCTGTAAATTTGTTAATTGATA |
Microarray Signals from CUST_22543_PI426222305
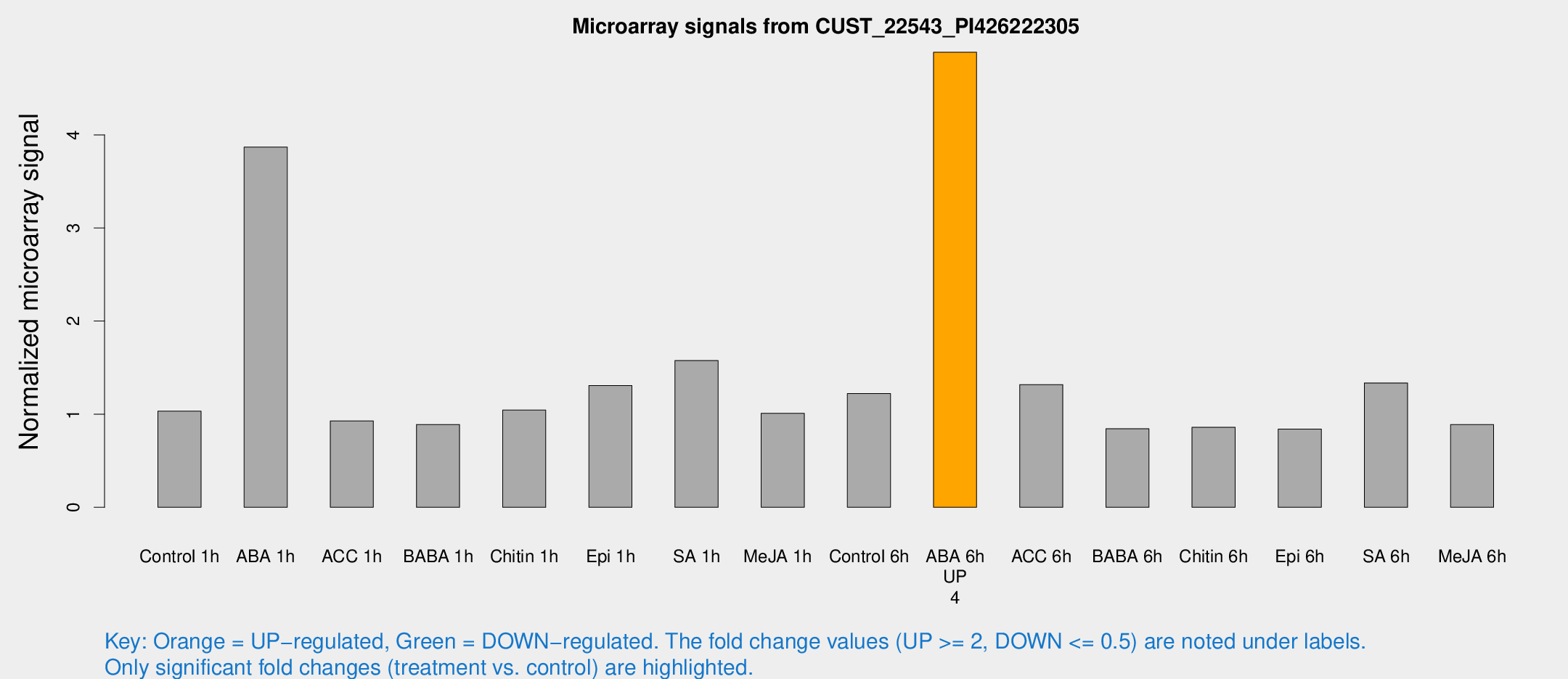
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.97626 | 2.98875 | 1.03263 | 0.499692 |
| ABA 1h | 24.3308 | 6.69876 | 3.86875 | 1.19959 |
| ACC 1h | 6.13492 | 3.40334 | 0.928172 | 0.512956 |
| BABA 1h | 5.49793 | 3.13284 | 0.889529 | 0.506098 |
| Chitin 1h | 6.06021 | 2.99525 | 1.04391 | 0.524452 |
| Epi 1h | 8.09133 | 2.97806 | 1.30825 | 0.612315 |
| SA 1h | 10.7597 | 3.0549 | 1.57569 | 0.486195 |
| Me-JA 1h | 5.25956 | 3.05609 | 1.00955 | 0.580483 |
| Control 6h | 9.29906 | 4.01869 | 1.22255 | 0.567253 |
| ABA 6h | 34.2606 | 5.83379 | 4.88744 | 0.724899 |
| ACC 6h | 9.87297 | 3.81359 | 1.31817 | 0.505405 |
| BABA 6h | 6.07842 | 3.43937 | 0.843678 | 0.477834 |
| Chitin 6h | 5.85622 | 3.3969 | 0.858672 | 0.497881 |
| Epi 6h | 6.10988 | 3.56849 | 0.839862 | 0.486893 |
| SA 6h | 9.07631 | 3.27489 | 1.33565 | 0.557602 |
| Me-JA 6h | 5.66895 | 3.08083 | 0.888242 | 0.482796 |
Source Transcript PGSC0003DMT400078034 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G58800.1 | +1 | 2e-101 | 304 | 161/202 (80%) | Quinone reductase family protein | chr5:23746032-23746895 REVERSE LENGTH=207 |
