Probe CUST_22489_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22489_PI426222305 | JHI_St_60k_v1 | DMT400039518 | ATTTCTGACTTGCAAGTGTATAAAATGCAAACAGTTCTCCGGAAACTATTCAAATGCTAG |
All Microarray Probes Designed to Gene DMG400015281
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22489_PI426222305 | JHI_St_60k_v1 | DMT400039518 | ATTTCTGACTTGCAAGTGTATAAAATGCAAACAGTTCTCCGGAAACTATTCAAATGCTAG |
| CUST_22496_PI426222305 | JHI_St_60k_v1 | DMT400039522 | ATTTCTGACTTGCAAGTGTATAAAATGCAAACAGTTCTCCGGAAACTATTCAAATGCTAG |
Microarray Signals from CUST_22489_PI426222305
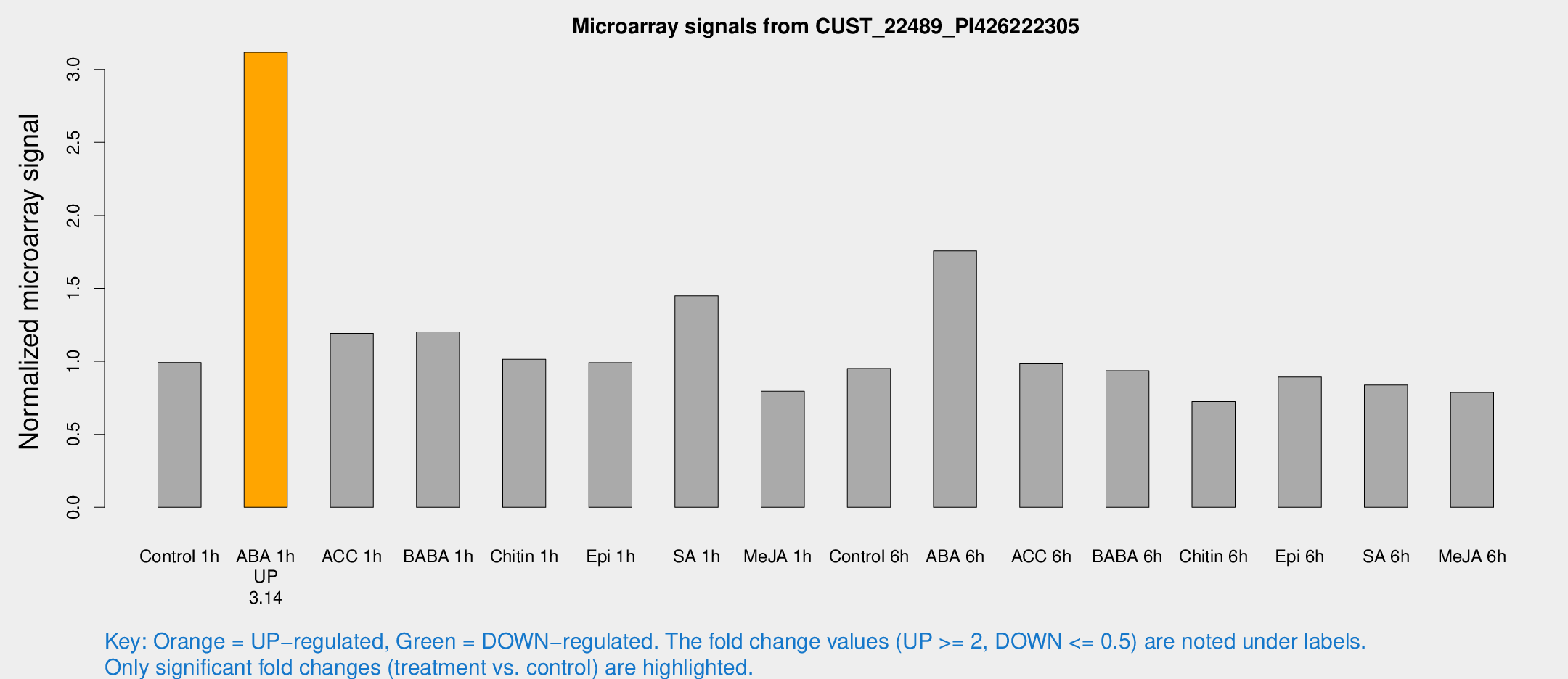
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 99.145 | 22.0049 | 0.992347 | 0.157824 |
| ABA 1h | 264.176 | 15.5759 | 3.11772 | 0.346563 |
| ACC 1h | 124.791 | 28.6543 | 1.19149 | 0.236271 |
| BABA 1h | 111.478 | 9.98983 | 1.20152 | 0.089006 |
| Chitin 1h | 90.7845 | 19.1994 | 1.01444 | 0.21284 |
| Epi 1h | 84.439 | 13.7417 | 0.991232 | 0.161055 |
| SA 1h | 142.429 | 8.8011 | 1.44874 | 0.108336 |
| Me-JA 1h | 62.8319 | 6.95893 | 0.795119 | 0.0613694 |
| Control 6h | 100.889 | 27.7741 | 0.951256 | 0.245554 |
| ABA 6h | 183.263 | 30.2441 | 1.75751 | 0.188403 |
| ACC 6h | 108.934 | 11.517 | 0.983521 | 0.0655897 |
| BABA 6h | 100.499 | 6.80491 | 0.935853 | 0.0697857 |
| Chitin 6h | 74.0593 | 5.52938 | 0.725155 | 0.054229 |
| Epi 6h | 96.8673 | 9.07087 | 0.891911 | 0.0856634 |
| SA 6h | 79.833 | 7.29916 | 0.837544 | 0.0604445 |
| Me-JA 6h | 79.1302 | 17.6714 | 0.787217 | 0.124789 |
Source Transcript PGSC0003DMT400039518 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G73390.1 | +2 | 1e-85 | 270 | 130/152 (86%) | Endosomal targeting BRO1-like domain-containing protein | chr1:27591079-27594148 REVERSE LENGTH=419 |
