Probe CUST_22416_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22416_PI426222305 | JHI_St_60k_v1 | DMT400039300 | GGGCAGGTGATGATGCTTTCCAAAACTTTACAAATTTCTTTTGCTTTGCAAATCTTGAAA |
All Microarray Probes Designed to Gene DMG401015201
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22416_PI426222305 | JHI_St_60k_v1 | DMT400039300 | GGGCAGGTGATGATGCTTTCCAAAACTTTACAAATTTCTTTTGCTTTGCAAATCTTGAAA |
Microarray Signals from CUST_22416_PI426222305
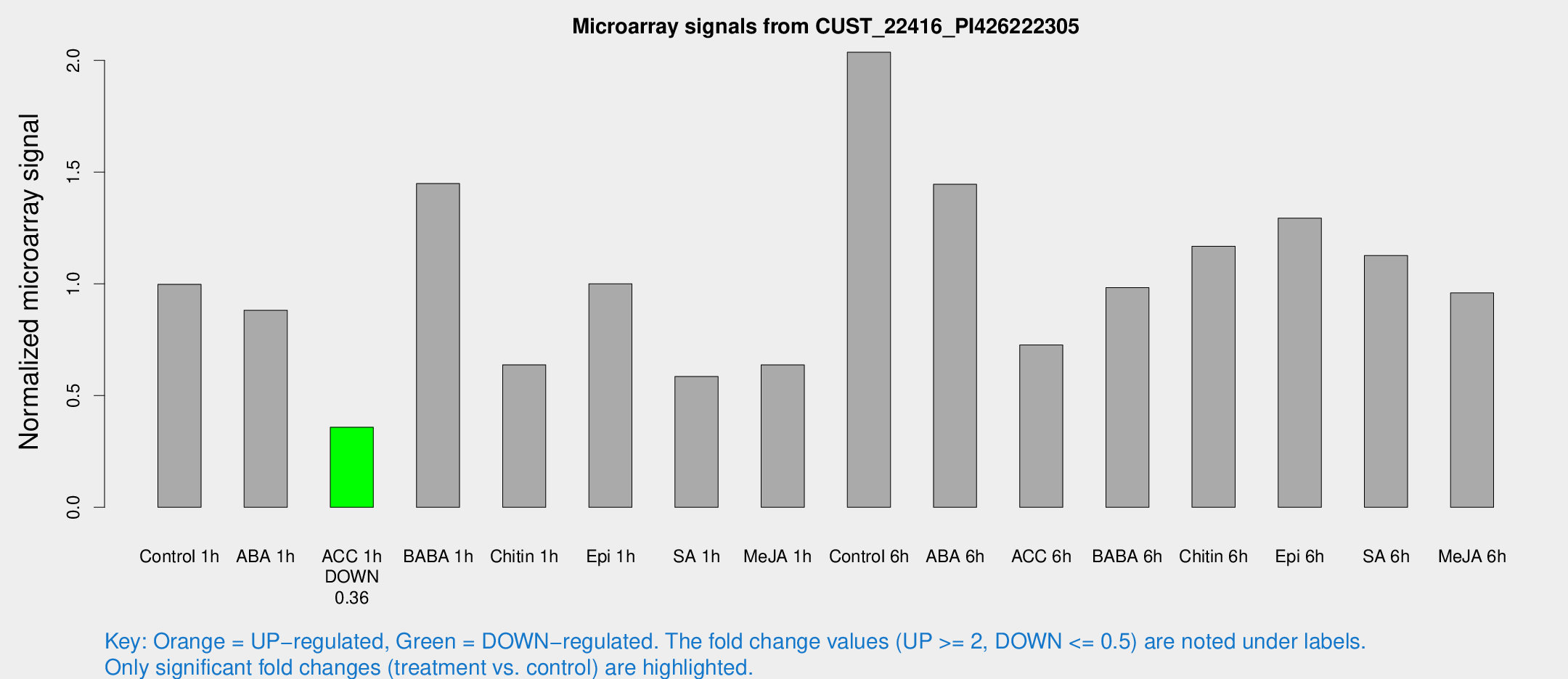
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 17.1084 | 3.1958 | 0.99781 | 0.194336 |
| ABA 1h | 15.221 | 5.46026 | 0.881996 | 0.349066 |
| ACC 1h | 6.3156 | 3.66508 | 0.358435 | 0.204313 |
| BABA 1h | 23.7214 | 3.5147 | 1.44873 | 0.216379 |
| Chitin 1h | 11.0567 | 3.88413 | 0.63766 | 0.233375 |
| Epi 1h | 16.2943 | 4.99534 | 1.0004 | 0.359869 |
| SA 1h | 10.1758 | 3.21393 | 0.584994 | 0.184706 |
| Me-JA 1h | 9.49899 | 3.17073 | 0.637181 | 0.247501 |
| Control 6h | 42.8799 | 18.4294 | 2.03645 | 0.951135 |
| ABA 6h | 35.4766 | 14.8429 | 1.44526 | 1.15691 |
| ACC 6h | 16.1037 | 5.06496 | 0.726405 | 0.231834 |
| BABA 6h | 25.1051 | 11.2551 | 0.983122 | 0.717929 |
| Chitin 6h | 23.3257 | 6.41162 | 1.16827 | 0.439159 |
| Epi 6h | 26.5369 | 6.78635 | 1.29415 | 0.450903 |
| SA 6h | 29.9862 | 18.421 | 1.12687 | 1.68572 |
| Me-JA 6h | 20.9362 | 8.24994 | 0.959511 | 0.569754 |
Source Transcript PGSC0003DMT400039300 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G36720.1 | +1 | 2e-22 | 87 | 41/79 (52%) | HVA22-like protein K | chr4:17307769-17309668 FORWARD LENGTH=200 |
