Probe CUST_22375_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22375_PI426222305 | JHI_St_60k_v1 | DMT400039411 | TAAACTACAAGTACCATGAGTTAAGAAAATCATTCTCGTTCCCTAACTGGTTGTCATCAC |
All Microarray Probes Designed to Gene DMG400015241
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22352_PI426222305 | JHI_St_60k_v1 | DMT400039410 | ACTACAGATCTATAACAGCAAAGAACTTCCTGACATCTGTTAGAAGGTATCAAGGGGAGC |
| CUST_22375_PI426222305 | JHI_St_60k_v1 | DMT400039411 | TAAACTACAAGTACCATGAGTTAAGAAAATCATTCTCGTTCCCTAACTGGTTGTCATCAC |
Microarray Signals from CUST_22375_PI426222305
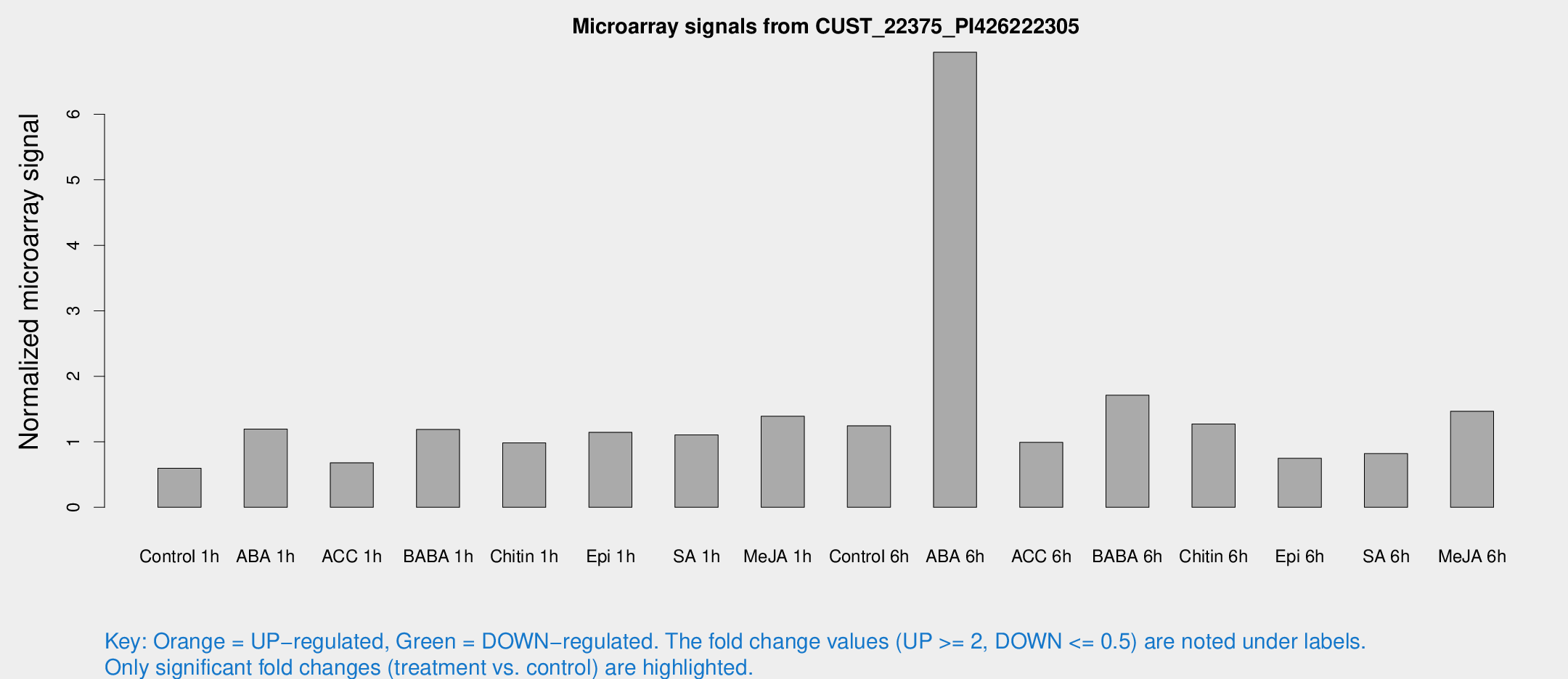
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.23068 | 3.03415 | 0.596172 | 0.344405 |
| ABA 1h | 10.3022 | 3.07524 | 1.19484 | 0.437015 |
| ACC 1h | 6.16974 | 3.3679 | 0.680163 | 0.372321 |
| BABA 1h | 11.422 | 4.25459 | 1.18853 | 0.447201 |
| Chitin 1h | 8.40092 | 3.06371 | 0.983918 | 0.417622 |
| Epi 1h | 9.589 | 3.15246 | 1.14469 | 0.43986 |
| SA 1h | 10.119 | 3.06699 | 1.1052 | 0.347158 |
| Me-JA 1h | 10.3946 | 3.12783 | 1.39055 | 0.472032 |
| Control 6h | 12.2293 | 3.4633 | 1.24452 | 0.435395 |
| ABA 6h | 76.6008 | 25.933 | 6.94769 | 3.05539 |
| ACC 6h | 10.9743 | 3.79427 | 0.992012 | 0.39818 |
| BABA 6h | 19.2509 | 6.31218 | 1.71097 | 0.645485 |
| Chitin 6h | 12.4666 | 3.48522 | 1.27262 | 0.405082 |
| Epi 6h | 7.70368 | 3.58981 | 0.748004 | 0.369081 |
| SA 6h | 7.23019 | 3.27052 | 0.820716 | 0.382252 |
| Me-JA 6h | 16.1082 | 7.5243 | 1.4664 | 0.918826 |
Source Transcript PGSC0003DMT400039411 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G48530.1 | +3 | 2e-114 | 353 | 215/318 (68%) | SNF1-related protein kinase regulatory subunit gamma 1 | chr3:17987559-17989592 FORWARD LENGTH=424 |
