Probe CUST_22211_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22211_PI426222305 | JHI_St_60k_v1 | DMT400047622 | GGGGTGTAGAATAGGTTGTATATTCCAACTCAAGTAGAAGTCAAGGAATTTTAATATGTG |
All Microarray Probes Designed to Gene DMG400018509
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22289_PI426222305 | JHI_St_60k_v1 | DMT400047621 | ATTCTGACTCTACTGCTACCGCGCAACTTATGGAAACGAATAAACTTTGTTTTGTTTTTT |
| CUST_22211_PI426222305 | JHI_St_60k_v1 | DMT400047622 | GGGGTGTAGAATAGGTTGTATATTCCAACTCAAGTAGAAGTCAAGGAATTTTAATATGTG |
Microarray Signals from CUST_22211_PI426222305
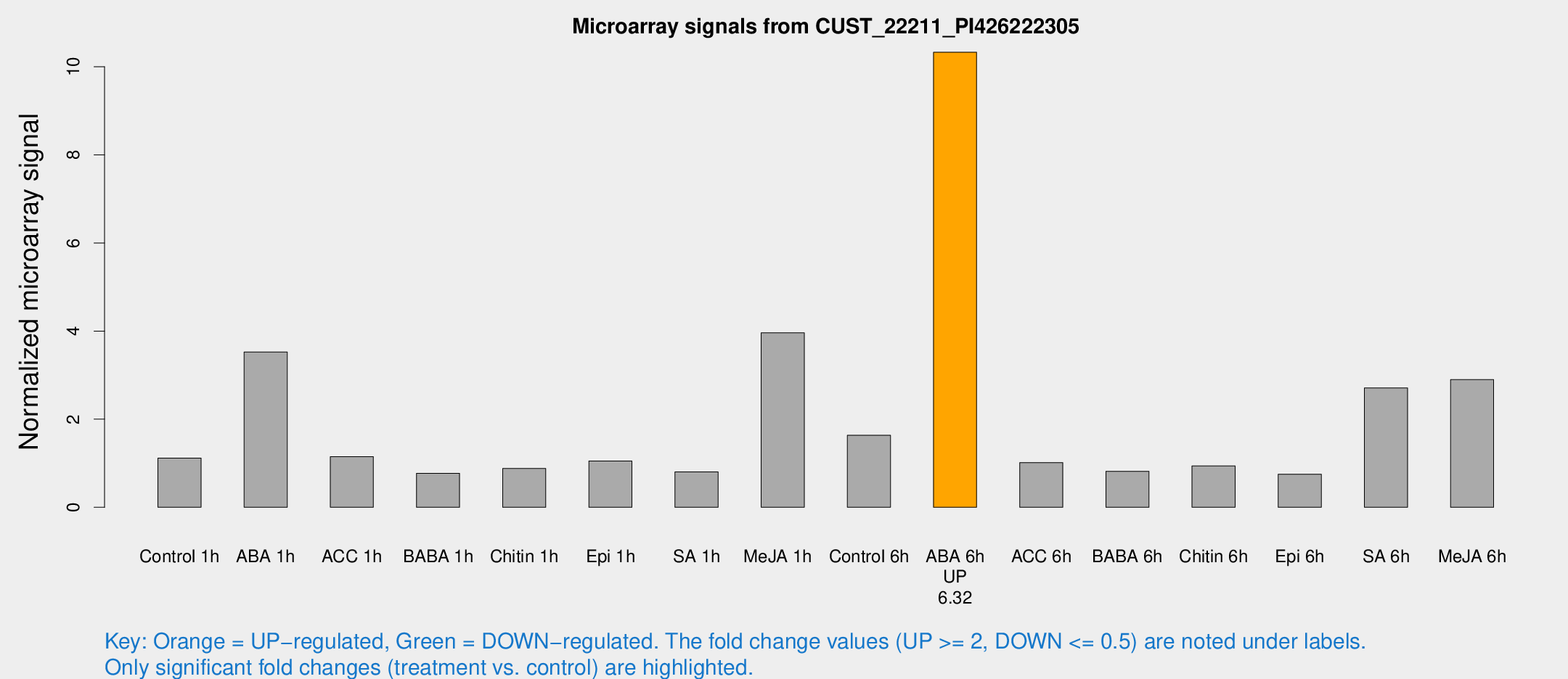
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.57573 | 3.27677 | 1.11665 | 0.466063 |
| ABA 1h | 26.4757 | 7.13779 | 3.52529 | 0.773323 |
| ACC 1h | 11.171 | 5.04198 | 1.14903 | 0.529664 |
| BABA 1h | 5.86745 | 3.4091 | 0.772753 | 0.448073 |
| Chitin 1h | 6.27694 | 3.29131 | 0.88033 | 0.464119 |
| Epi 1h | 7.8031 | 3.21243 | 1.05249 | 0.506438 |
| SA 1h | 6.65705 | 3.18734 | 0.802447 | 0.405752 |
| Me-JA 1h | 25.7461 | 3.77493 | 3.961 | 0.585776 |
| Control 6h | 17.6457 | 10.0195 | 1.63485 | 0.999872 |
| ABA 6h | 93.0519 | 22.5056 | 10.3298 | 2.61541 |
| ACC 6h | 10.4356 | 4.16285 | 1.01462 | 0.463829 |
| BABA 6h | 7.23361 | 3.85663 | 0.817506 | 0.438662 |
| Chitin 6h | 8.2166 | 3.83041 | 0.939397 | 0.472671 |
| Epi 6h | 6.73385 | 3.94807 | 0.750789 | 0.434875 |
| SA 6h | 23.2917 | 7.36146 | 2.70917 | 1.33867 |
| Me-JA 6h | 22.7654 | 3.75905 | 2.89851 | 0.485493 |
Source Transcript PGSC0003DMT400047622 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G46680.1 | +1 | 5e-42 | 149 | 93/215 (43%) | homeobox 7 | chr2:19165777-19166773 REVERSE LENGTH=258 |
