Probe CUST_21815_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21815_PI426222305 | JHI_St_60k_v1 | DMT400024887 | GCTGCCAGAAAAGATGACCTTATTTTGGTTGCCACTCCTTATCAACCACCTCAACAATGA |
All Microarray Probes Designed to Gene DMG400009621
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21815_PI426222305 | JHI_St_60k_v1 | DMT400024887 | GCTGCCAGAAAAGATGACCTTATTTTGGTTGCCACTCCTTATCAACCACCTCAACAATGA |
Microarray Signals from CUST_21815_PI426222305
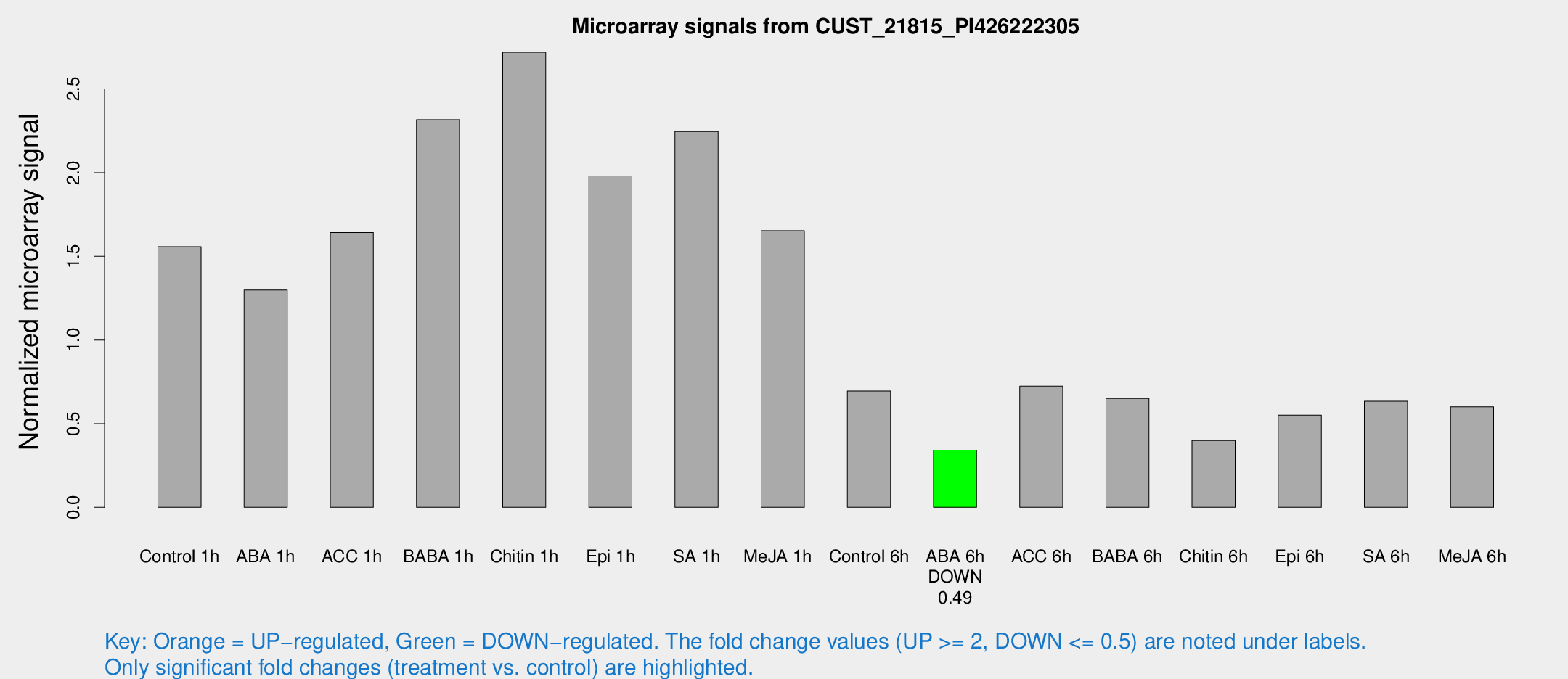
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1820.92 | 122.621 | 1.55742 | 0.183053 |
| ABA 1h | 1342.63 | 84.7065 | 1.29866 | 0.135398 |
| ACC 1h | 2007.86 | 288.556 | 1.6421 | 0.123816 |
| BABA 1h | 2821.18 | 717.286 | 2.31633 | 0.47257 |
| Chitin 1h | 2864.42 | 233.819 | 2.71931 | 0.161201 |
| Epi 1h | 2039.92 | 303.523 | 1.98074 | 0.297099 |
| SA 1h | 2713.5 | 265.996 | 2.24607 | 0.12971 |
| Me-JA 1h | 1595.3 | 198.743 | 1.65326 | 0.0955276 |
| Control 6h | 835.732 | 136.566 | 0.694883 | 0.0644944 |
| ABA 6h | 436.812 | 77.9139 | 0.342083 | 0.0418099 |
| ACC 6h | 969.288 | 56.2717 | 0.724165 | 0.0856399 |
| BABA 6h | 851.435 | 56.3944 | 0.650501 | 0.0376939 |
| Chitin 6h | 496.427 | 31.3286 | 0.398999 | 0.0232755 |
| Epi 6h | 738.389 | 101.765 | 0.551235 | 0.0555685 |
| SA 6h | 750.768 | 128.286 | 0.634297 | 0.113704 |
| Me-JA 6h | 715.026 | 110.571 | 0.600178 | 0.059336 |
Source Transcript PGSC0003DMT400024887 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G26310.1 | +1 | 1e-123 | 374 | 203/458 (44%) | cytochrome P450, family 71, subfamily B, polypeptide 35 | chr3:9641089-9642779 REVERSE LENGTH=500 |
