Probe CUST_21509_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21509_PI426222305 | JHI_St_60k_v1 | DMT400070038 | GACACGTGTTAGACAATAAATTTTCAGAATATCTTTAGTATCCGCCAAAATTTTGTGCGC |
All Microarray Probes Designed to Gene DMG400027238
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21421_PI426222305 | JHI_St_60k_v1 | DMT400070037 | TGCTGTGATTAATGGATGGGAAAATGAAGAAGTTCAAAAGGCTACTTCATGGATGAAGAA |
| CUST_21509_PI426222305 | JHI_St_60k_v1 | DMT400070038 | GACACGTGTTAGACAATAAATTTTCAGAATATCTTTAGTATCCGCCAAAATTTTGTGCGC |
Microarray Signals from CUST_21509_PI426222305
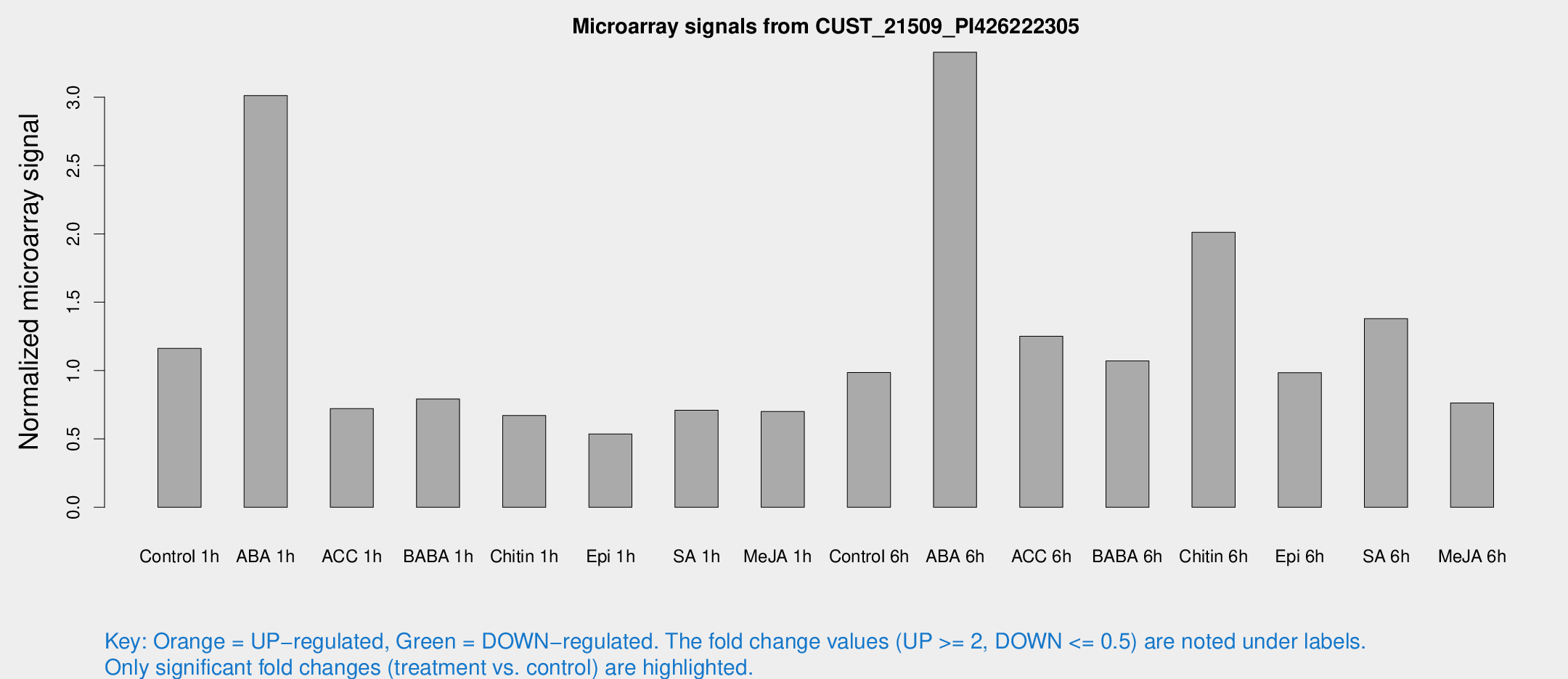
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 17.4862 | 3.28264 | 1.16227 | 0.225979 |
| ABA 1h | 40.7286 | 7.74911 | 3.0119 | 0.557818 |
| ACC 1h | 12.4934 | 4.61422 | 0.721983 | 0.275875 |
| BABA 1h | 13.2118 | 5.06736 | 0.79181 | 0.286601 |
| Chitin 1h | 9.50977 | 3.15441 | 0.671174 | 0.263853 |
| Epi 1h | 7.29544 | 3.10817 | 0.536448 | 0.257719 |
| SA 1h | 10.7877 | 3.14399 | 0.710549 | 0.207118 |
| Me-JA 1h | 8.97723 | 3.19875 | 0.700943 | 0.280885 |
| Control 6h | 18.3345 | 7.71869 | 0.98571 | 0.487637 |
| ABA 6h | 56.8784 | 14.5181 | 3.32852 | 0.907436 |
| ACC 6h | 21.2435 | 4.05683 | 1.25081 | 0.23584 |
| BABA 6h | 18.2896 | 3.68698 | 1.07068 | 0.2325 |
| Chitin 6h | 32.8901 | 6.15127 | 2.01092 | 0.40494 |
| Epi 6h | 16.8517 | 3.79673 | 0.984174 | 0.229826 |
| SA 6h | 22.953 | 6.87944 | 1.3804 | 0.761231 |
| Me-JA 6h | 19.9942 | 14.6385 | 0.762643 | 0.747102 |
Source Transcript PGSC0003DMT400070038 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G57540.1 | +1 | 4e-06 | 47 | 22/27 (81%) | Remorin family protein | chr3:21301623-21302924 REVERSE LENGTH=296 |
