Probe CUST_21473_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21473_PI426222305 | JHI_St_60k_v1 | DMT400070218 | CGCGTATATCACAAGATAGACTTGCTTTTTATGGCTATTAGTAAGATGATTGCTATGTAC |
All Microarray Probes Designed to Gene DMG400027291
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21473_PI426222305 | JHI_St_60k_v1 | DMT400070218 | CGCGTATATCACAAGATAGACTTGCTTTTTATGGCTATTAGTAAGATGATTGCTATGTAC |
Microarray Signals from CUST_21473_PI426222305
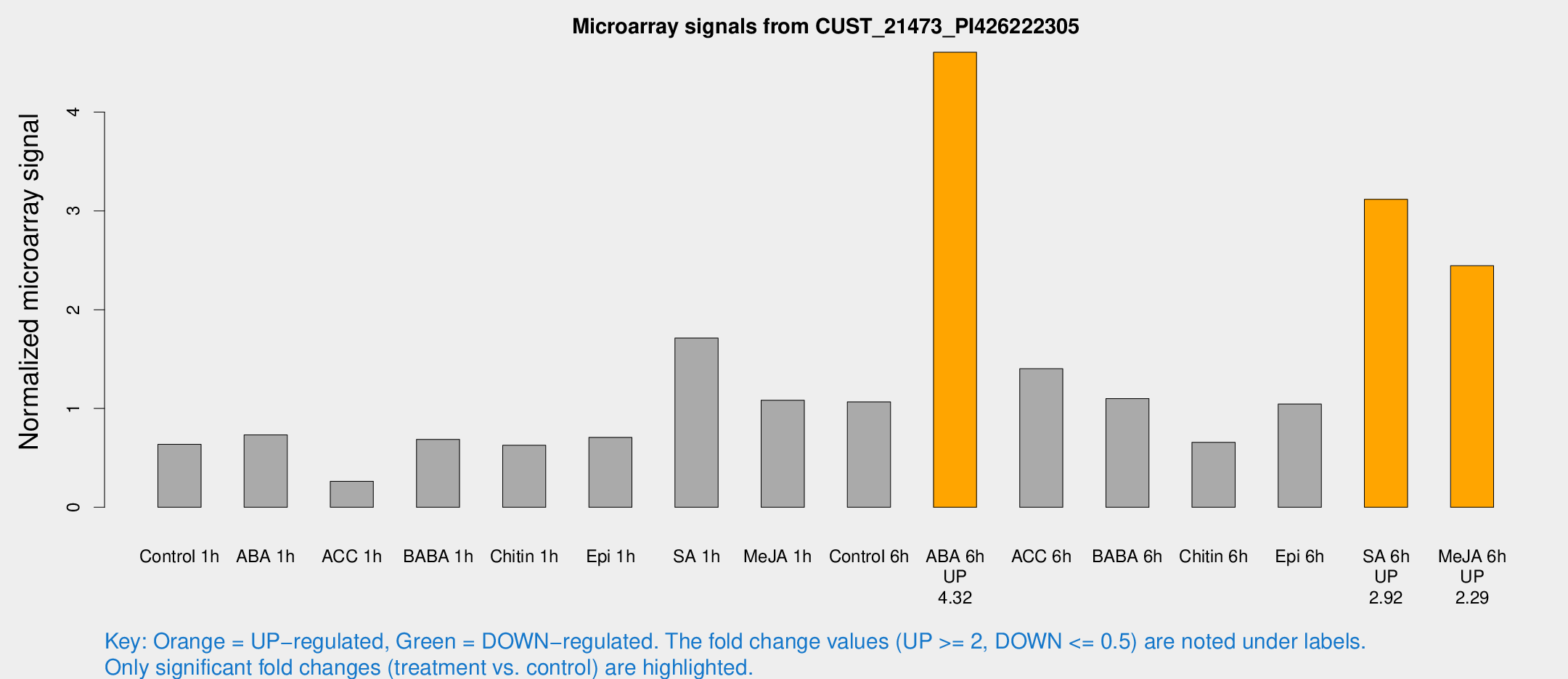
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 770.11 | 133.609 | 0.637555 | 0.163379 |
| ABA 1h | 829.426 | 220.558 | 0.733017 | 0.191544 |
| ACC 1h | 441.538 | 206.027 | 0.263227 | 0.195583 |
| BABA 1h | 882.58 | 264.221 | 0.687711 | 0.206394 |
| Chitin 1h | 709.393 | 175.02 | 0.628078 | 0.172132 |
| Epi 1h | 755.279 | 164.377 | 0.707388 | 0.167013 |
| SA 1h | 2277.9 | 655.942 | 1.71209 | 0.493109 |
| Me-JA 1h | 1080.51 | 220.843 | 1.08422 | 0.128231 |
| Control 6h | 1278.43 | 180.872 | 1.06608 | 0.0799671 |
| ABA 6h | 6128.96 | 1503.98 | 4.60525 | 1.00472 |
| ACC 6h | 1883.89 | 108.873 | 1.40232 | 0.210085 |
| BABA 6h | 1469.59 | 202.698 | 1.10046 | 0.108947 |
| Chitin 6h | 820.96 | 47.7018 | 0.65707 | 0.0380839 |
| Epi 6h | 1424.33 | 241.618 | 1.04461 | 0.1155 |
| SA 6h | 3743.13 | 686.744 | 3.11725 | 0.301516 |
| Me-JA 6h | 2862.53 | 188.319 | 2.44483 | 0.149935 |
Source Transcript PGSC0003DMT400070218 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G15490.1 | +1 | 5e-50 | 181 | 114/334 (34%) | UDP-glycosyltransferase 73B4 | chr2:6761750-6763398 FORWARD LENGTH=484 |
