Probe CUST_21421_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21421_PI426222305 | JHI_St_60k_v1 | DMT400070037 | TGCTGTGATTAATGGATGGGAAAATGAAGAAGTTCAAAAGGCTACTTCATGGATGAAGAA |
All Microarray Probes Designed to Gene DMG400027238
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21421_PI426222305 | JHI_St_60k_v1 | DMT400070037 | TGCTGTGATTAATGGATGGGAAAATGAAGAAGTTCAAAAGGCTACTTCATGGATGAAGAA |
| CUST_21509_PI426222305 | JHI_St_60k_v1 | DMT400070038 | GACACGTGTTAGACAATAAATTTTCAGAATATCTTTAGTATCCGCCAAAATTTTGTGCGC |
Microarray Signals from CUST_21421_PI426222305
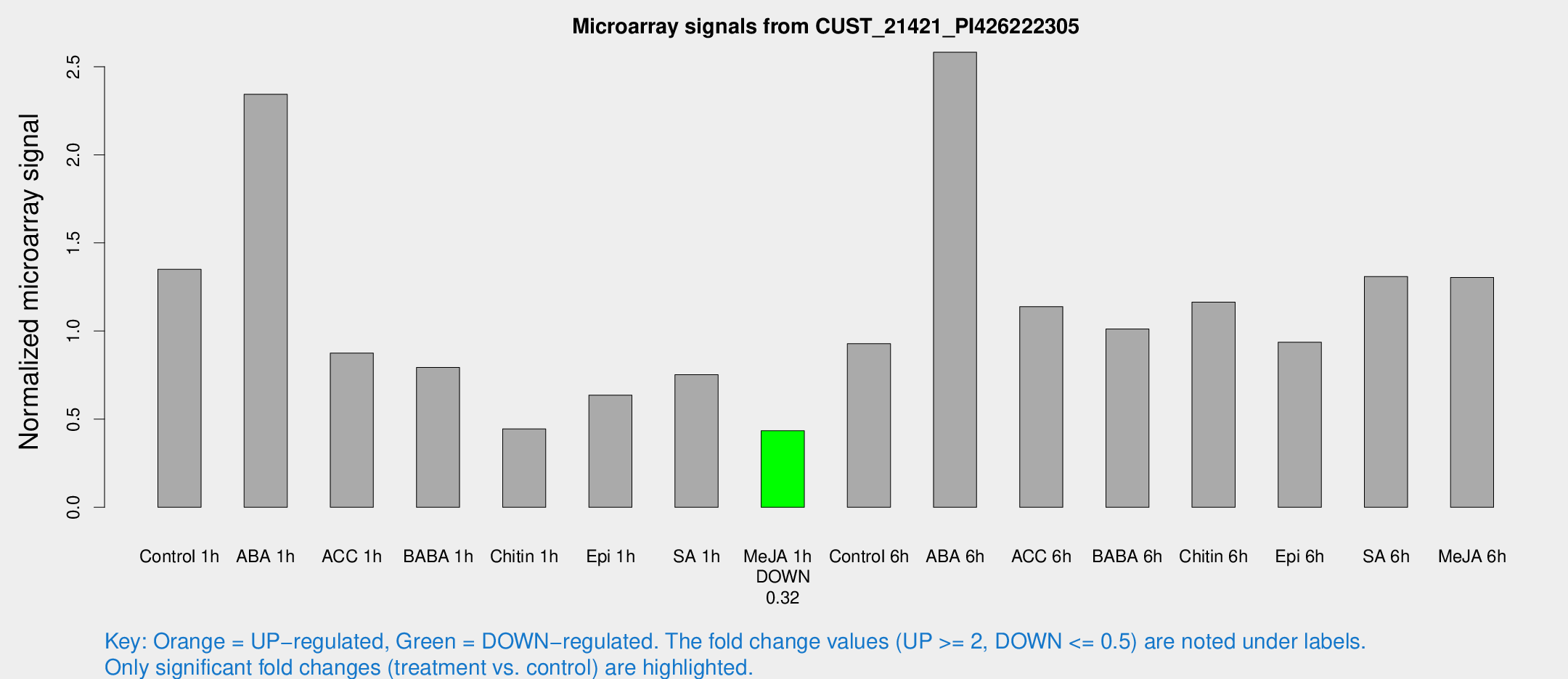
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1874.17 | 193.657 | 1.35003 | 0.0779921 |
| ABA 1h | 2971.36 | 582.173 | 2.343 | 0.424038 |
| ACC 1h | 1433 | 501.734 | 0.875135 | 0.32327 |
| BABA 1h | 1104.79 | 232.83 | 0.793419 | 0.10843 |
| Chitin 1h | 651.779 | 230.781 | 0.444931 | 0.223416 |
| Epi 1h | 913.891 | 317.53 | 0.636679 | 0.354157 |
| SA 1h | 1098.97 | 212.287 | 0.752779 | 0.0959065 |
| Me-JA 1h | 505.013 | 101.128 | 0.434435 | 0.0501698 |
| Control 6h | 1425 | 456.217 | 0.927916 | 0.261708 |
| ABA 6h | 3855.51 | 559.224 | 2.5819 | 0.272624 |
| ACC 6h | 1865.3 | 343.823 | 1.13877 | 0.214386 |
| BABA 6h | 1574.25 | 181.676 | 1.01148 | 0.0992797 |
| Chitin 6h | 1716.89 | 169.771 | 1.16368 | 0.138201 |
| Epi 6h | 1504.73 | 292.764 | 0.93624 | 0.112917 |
| SA 6h | 1818.42 | 258.675 | 1.30953 | 0.0756709 |
| Me-JA 6h | 1818.15 | 264.175 | 1.30408 | 0.0931267 |
Source Transcript PGSC0003DMT400070037 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G41870.1 | +2 | 1e-38 | 144 | 128/224 (57%) | Remorin family protein | chr2:17471119-17472519 REVERSE LENGTH=274 |
