Probe CUST_19764_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19764_PI426222305 | JHI_St_60k_v1 | DMT400064111 | GGGAGACACTCCTACTTGTATCTTGAGAATGTTTAATCTTTGGCATTTTATCTTTCTCTT |
All Microarray Probes Designed to Gene DMG400024912
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19764_PI426222305 | JHI_St_60k_v1 | DMT400064111 | GGGAGACACTCCTACTTGTATCTTGAGAATGTTTAATCTTTGGCATTTTATCTTTCTCTT |
Microarray Signals from CUST_19764_PI426222305
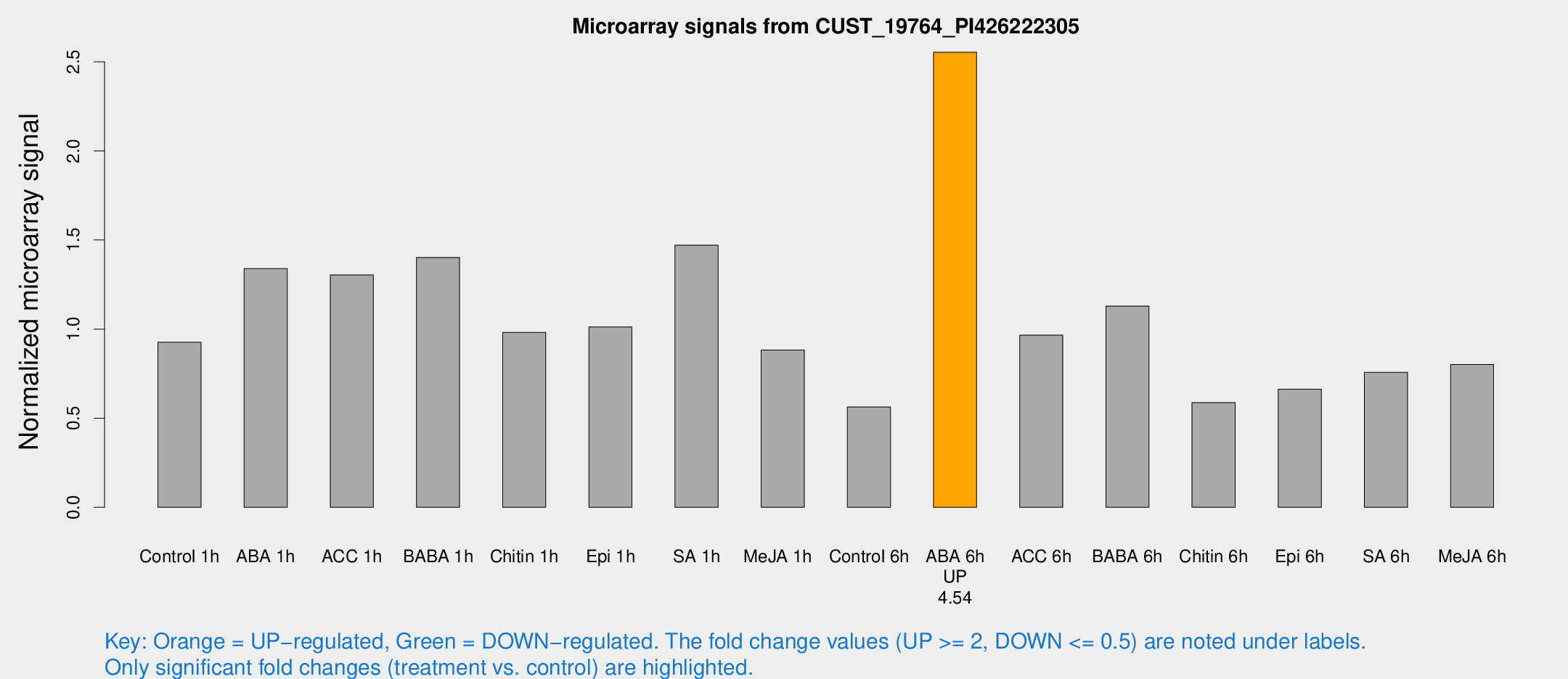
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 801.632 | 142.733 | 0.92598 | 0.132549 |
| ABA 1h | 1027.03 | 169.618 | 1.33979 | 0.177203 |
| ACC 1h | 1190.03 | 261.768 | 1.30372 | 0.234184 |
| BABA 1h | 1134.63 | 65.6038 | 1.40205 | 0.1379 |
| Chitin 1h | 757.622 | 109.591 | 0.981566 | 0.143403 |
| Epi 1h | 766.482 | 147.708 | 1.01231 | 0.208773 |
| SA 1h | 1314.78 | 259.246 | 1.47124 | 0.201759 |
| Me-JA 1h | 608.058 | 45.0522 | 0.882385 | 0.0666398 |
| Control 6h | 558.247 | 187.039 | 0.562733 | 0.214047 |
| ABA 6h | 2304.19 | 247.801 | 2.55344 | 0.172373 |
| ACC 6h | 948.412 | 136.162 | 0.965718 | 0.0559077 |
| BABA 6h | 1120.4 | 257.307 | 1.12894 | 0.266257 |
| Chitin 6h | 535.597 | 78.6125 | 0.587314 | 0.0827456 |
| Epi 6h | 647.965 | 111.224 | 0.662862 | 0.0914218 |
| SA 6h | 629.835 | 36.601 | 0.757104 | 0.0603901 |
| Me-JA 6h | 687.322 | 109.801 | 0.800905 | 0.103509 |
Source Transcript PGSC0003DMT400064111 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G24275.1 | +2 | 6e-07 | 49 | 32/67 (48%) | Identified as a screen for stress-responsive genes. | chr4:12588579-12588959 FORWARD LENGTH=126 |
