Probe CUST_19451_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19451_PI426222305 | JHI_St_60k_v1 | DMT400072823 | CAAAGGCTCTTCAAGAAAAAGGTTGGTCTCAAGAGATTATGGATATTTATCTTAGAGAAC |
All Microarray Probes Designed to Gene DMG400028330
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19451_PI426222305 | JHI_St_60k_v1 | DMT400072823 | CAAAGGCTCTTCAAGAAAAAGGTTGGTCTCAAGAGATTATGGATATTTATCTTAGAGAAC |
Microarray Signals from CUST_19451_PI426222305
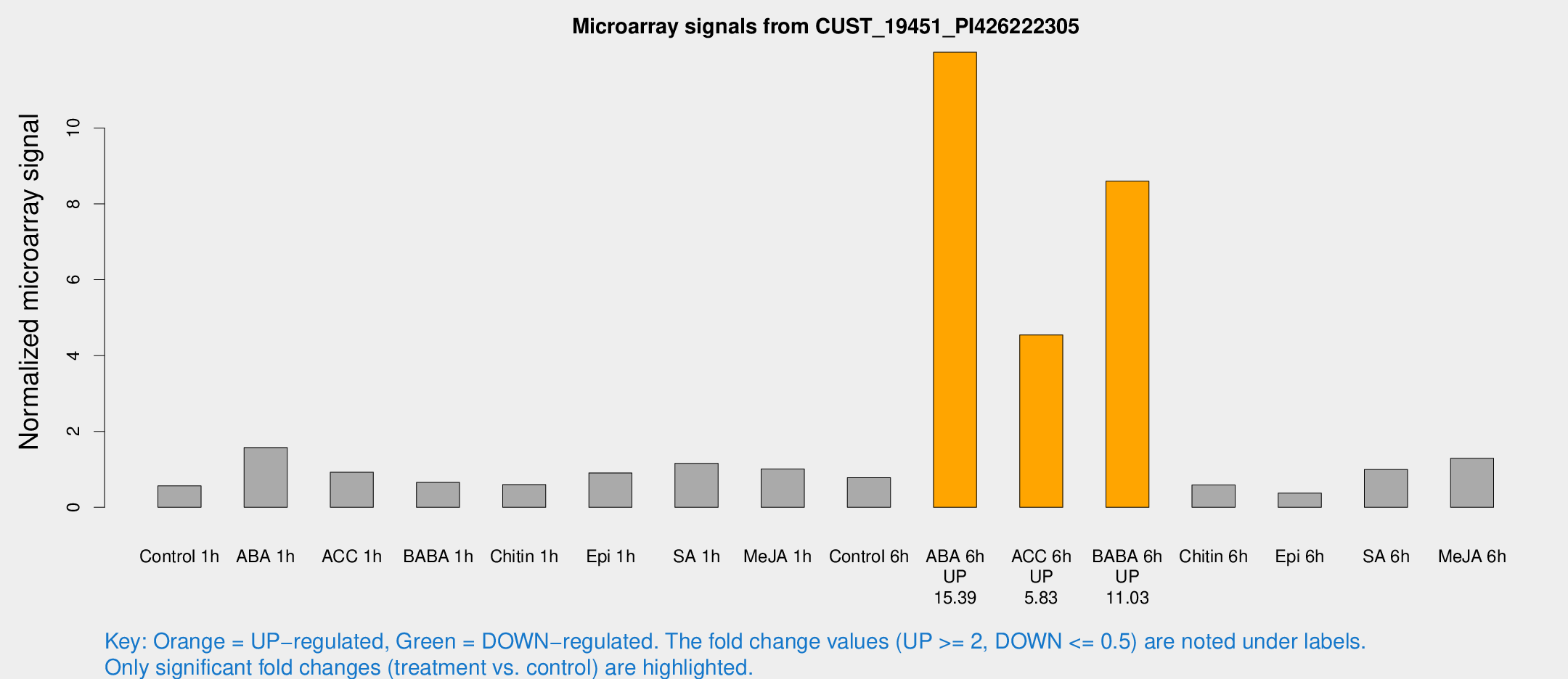
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 10.6131 | 3.14206 | 0.567532 | 0.208498 |
| ABA 1h | 24.326 | 4.32984 | 1.57664 | 0.415024 |
| ACC 1h | 16.1592 | 3.6251 | 0.923503 | 0.212031 |
| BABA 1h | 11.9892 | 3.53491 | 0.657166 | 0.237107 |
| Chitin 1h | 10.0742 | 3.42789 | 0.597477 | 0.219533 |
| Epi 1h | 13.7227 | 3.11006 | 0.90636 | 0.230141 |
| SA 1h | 21.3859 | 4.78436 | 1.15792 | 0.249293 |
| Me-JA 1h | 14.0698 | 3.23238 | 1.00902 | 0.234798 |
| Control 6h | 15.574 | 5.27051 | 0.779729 | 0.296824 |
| ABA 6h | 246.727 | 78.6611 | 11.9982 | 4.50715 |
| ACC 6h | 88.9974 | 9.31101 | 4.54599 | 0.93427 |
| BABA 6h | 184.764 | 64.8955 | 8.59673 | 3.30393 |
| Chitin 6h | 11.055 | 3.52994 | 0.59075 | 0.207351 |
| Epi 6h | 7.46978 | 3.67598 | 0.376829 | 0.192793 |
| SA 6h | 17.7535 | 4.05521 | 0.994983 | 0.237204 |
| Me-JA 6h | 23.3125 | 6.38302 | 1.29173 | 0.296451 |
Source Transcript PGSC0003DMT400072823 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G03360.2 | +2 | 2e-131 | 393 | 207/420 (49%) | Glycosyltransferase family 61 protein | chr2:1022287-1024273 REVERSE LENGTH=455 |
