Probe CUST_19416_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19416_PI426222305 | JHI_St_60k_v1 | DMT400072921 | TAAGCAGCCTACCTTGATGCAATTGTTAAAGACAGAATCCATGGCTCAAACCCGTGAGCT |
All Microarray Probes Designed to Gene DMG400028369
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19340_PI426222305 | JHI_St_60k_v1 | DMT400072922 | CATGTCACCATCATCATCCATTAGATGTTCTATTTCATCACATACCTGAACATTTTTCGA |
| CUST_19416_PI426222305 | JHI_St_60k_v1 | DMT400072921 | TAAGCAGCCTACCTTGATGCAATTGTTAAAGACAGAATCCATGGCTCAAACCCGTGAGCT |
Microarray Signals from CUST_19416_PI426222305
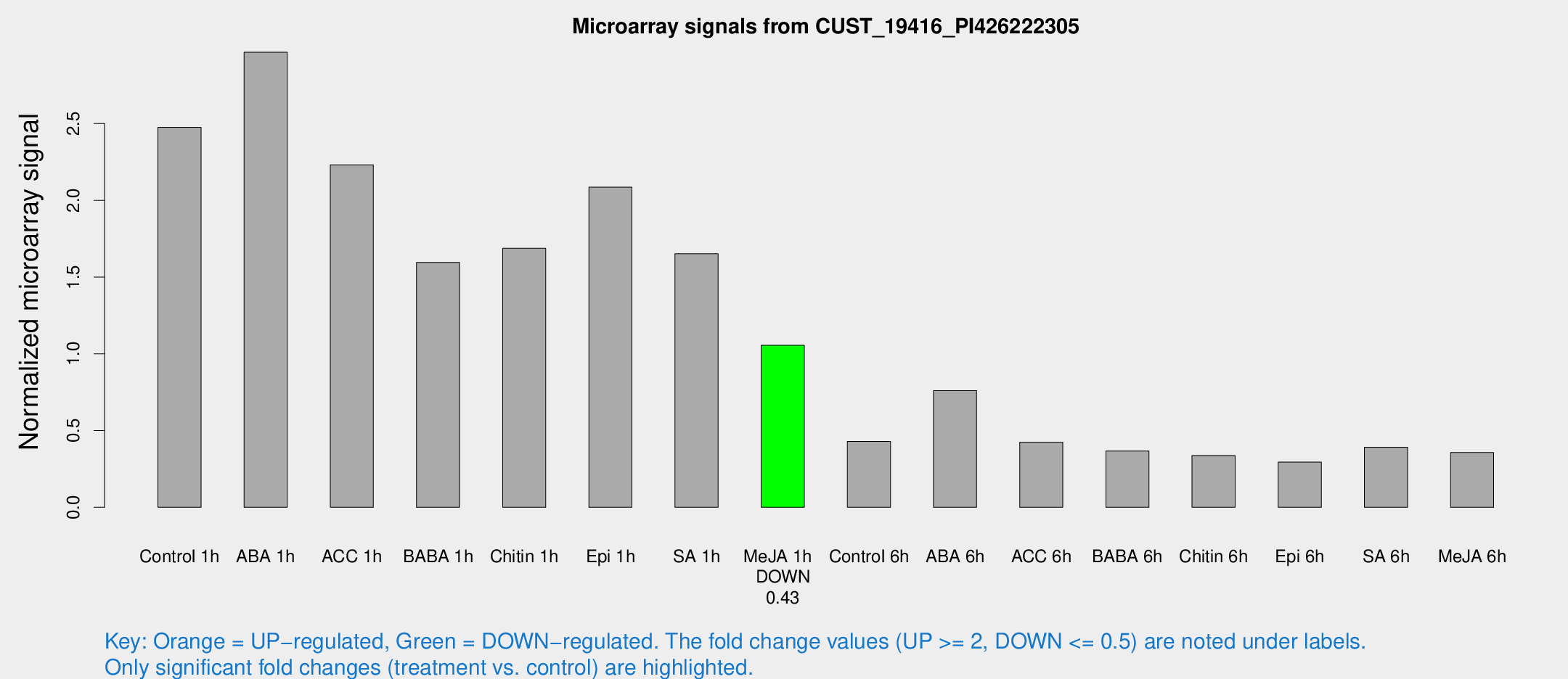
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 664.752 | 80.2358 | 2.47506 | 0.14334 |
| ABA 1h | 697.267 | 43.6821 | 2.96474 | 0.38272 |
| ACC 1h | 610.936 | 52.8719 | 2.2307 | 0.187123 |
| BABA 1h | 453.414 | 126.674 | 1.59575 | 0.410392 |
| Chitin 1h | 406.834 | 46.5106 | 1.687 | 0.188985 |
| Epi 1h | 477.963 | 27.7495 | 2.08533 | 0.121058 |
| SA 1h | 472.872 | 111.604 | 1.65174 | 0.277796 |
| Me-JA 1h | 230.291 | 21.1696 | 1.05591 | 0.0625533 |
| Control 6h | 128.403 | 46.3245 | 0.429622 | 0.136815 |
| ABA 6h | 228.373 | 53.7156 | 0.759653 | 0.18456 |
| ACC 6h | 132.569 | 22.7814 | 0.424699 | 0.0271252 |
| BABA 6h | 109.668 | 10.3702 | 0.366722 | 0.0278997 |
| Chitin 6h | 102.502 | 28.698 | 0.337519 | 0.087684 |
| Epi 6h | 92.1783 | 18.5828 | 0.294483 | 0.0450925 |
| SA 6h | 125.091 | 51.9599 | 0.39114 | 0.154973 |
| Me-JA 6h | 94.4012 | 6.24941 | 0.356676 | 0.0235874 |
Source Transcript PGSC0003DMT400072921 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G60980.2 | +1 | 4e-31 | 126 | 101/320 (32%) | Nuclear transport factor 2 (NTF2) family protein with RNA binding (RRM-RBD-RNP motifs) domain | chr5:24543988-24546027 FORWARD LENGTH=460 |
